Academic Figures: Common Pitfalls and Best Practices
Things that annoy me - An opinionated guide
Jan Stanstrup
Introduction
Why This Matters
Figures are often the first (and sometimes only) thing readers look at
Poor quality figures look unprofessional
Low resolution images look bad when scaled up (e.g. posters)
Bad visualizations can be misleading
What We’ll Cover
Color Gradients
What is the Rainbow Scale?
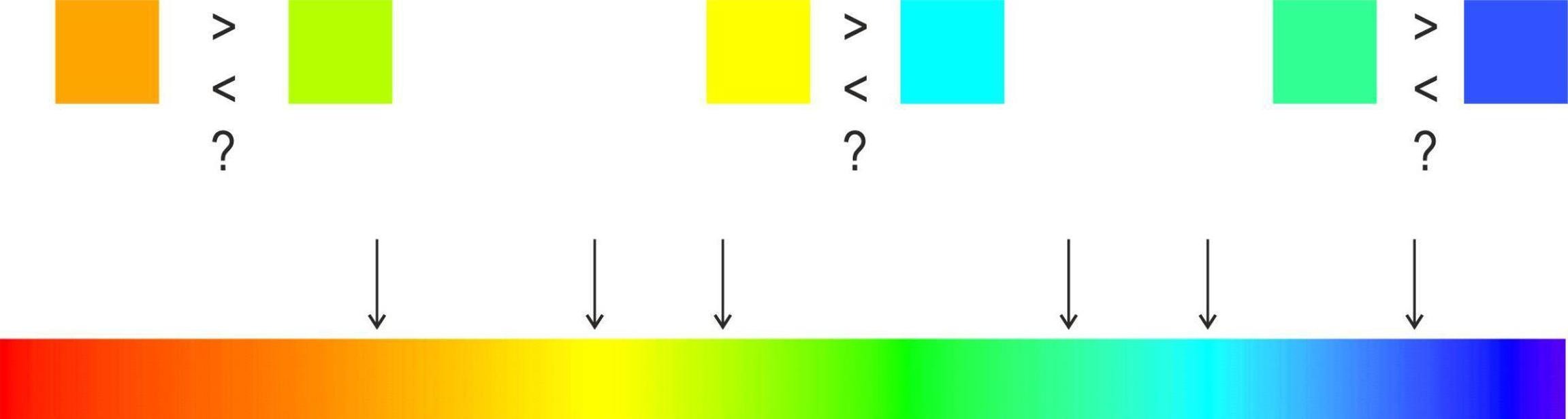
Interactive Challenge: Can you order these colors?

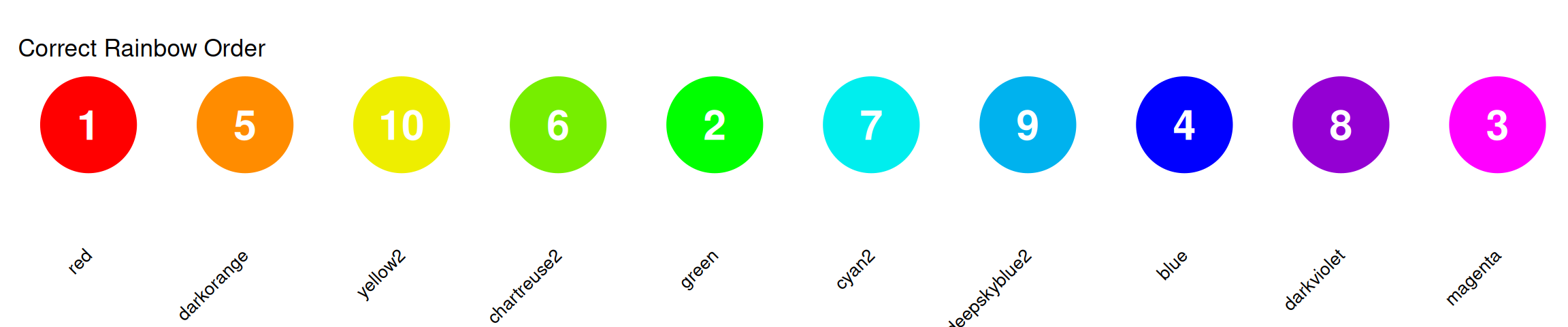
The Correct Rainbow Order
Colors in order: Red → Orange → Yellow → Green → Cyan → Blue → Purple/Magenta
Comparing Rainbow Implementations
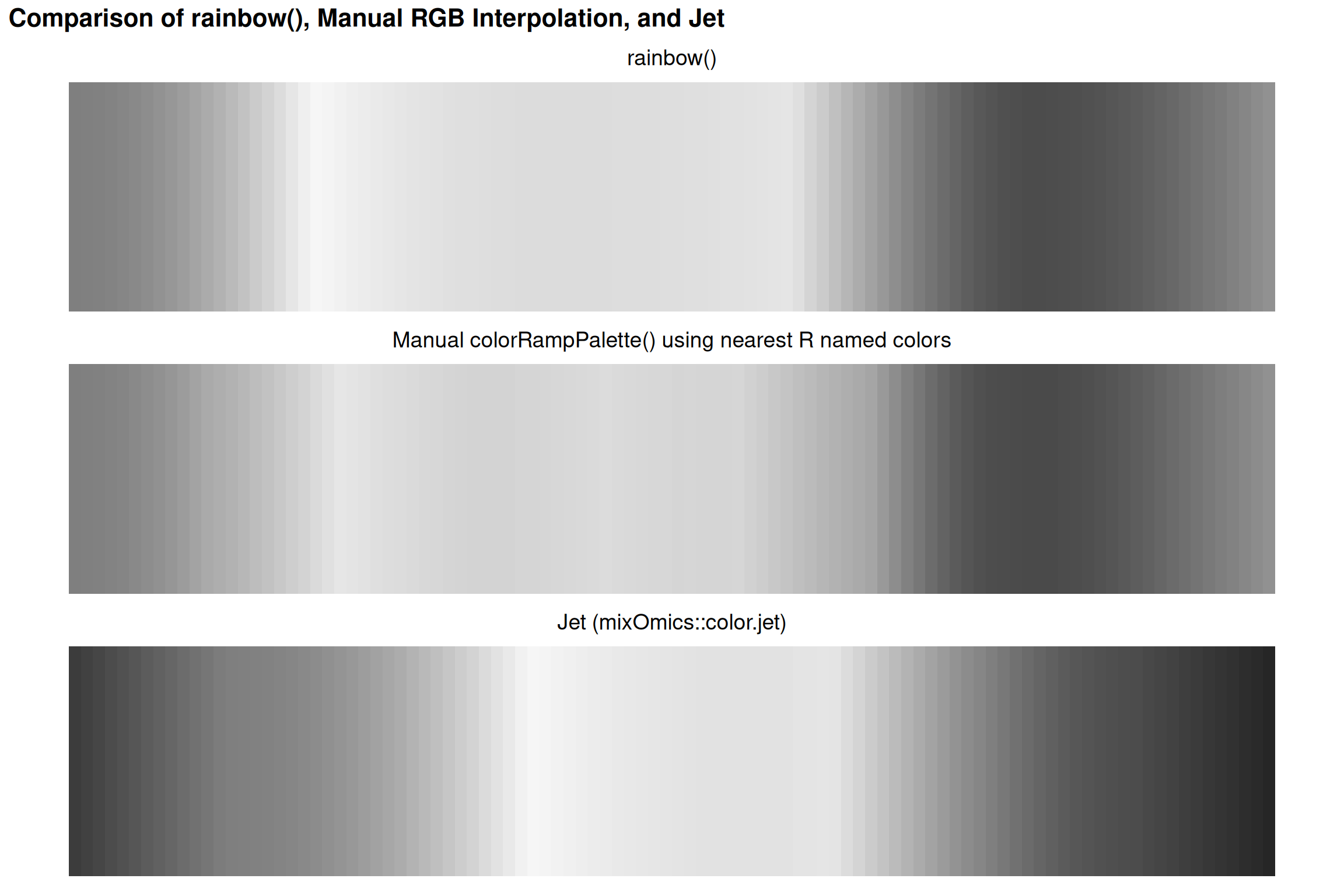


Why This Order?
This follows the visible light spectrum by wavelength:
- Red: ~700 nm (longest)
- Violet: ~400 nm (shortest)
But wavelength order ≠ perceptual order!
A Tale of Two Colormaps
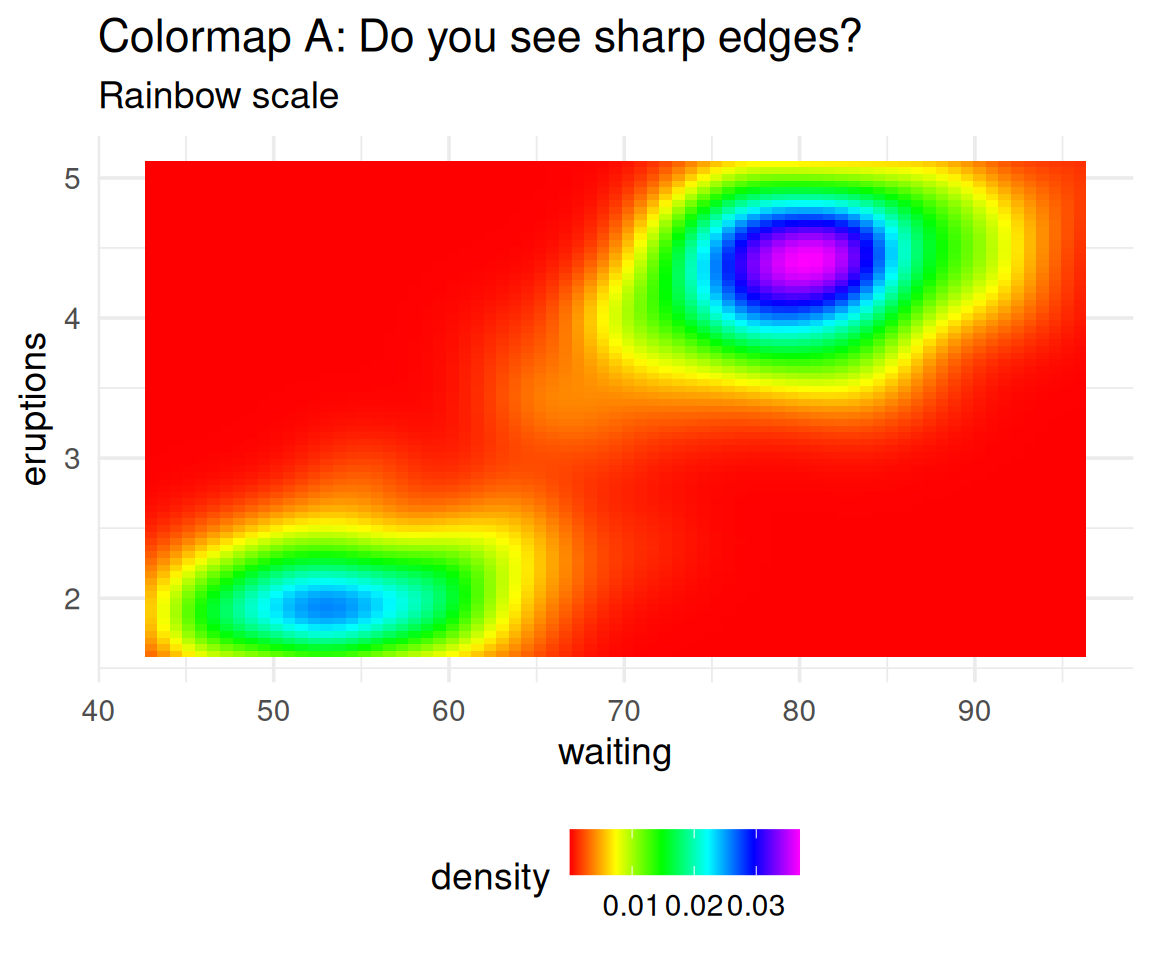
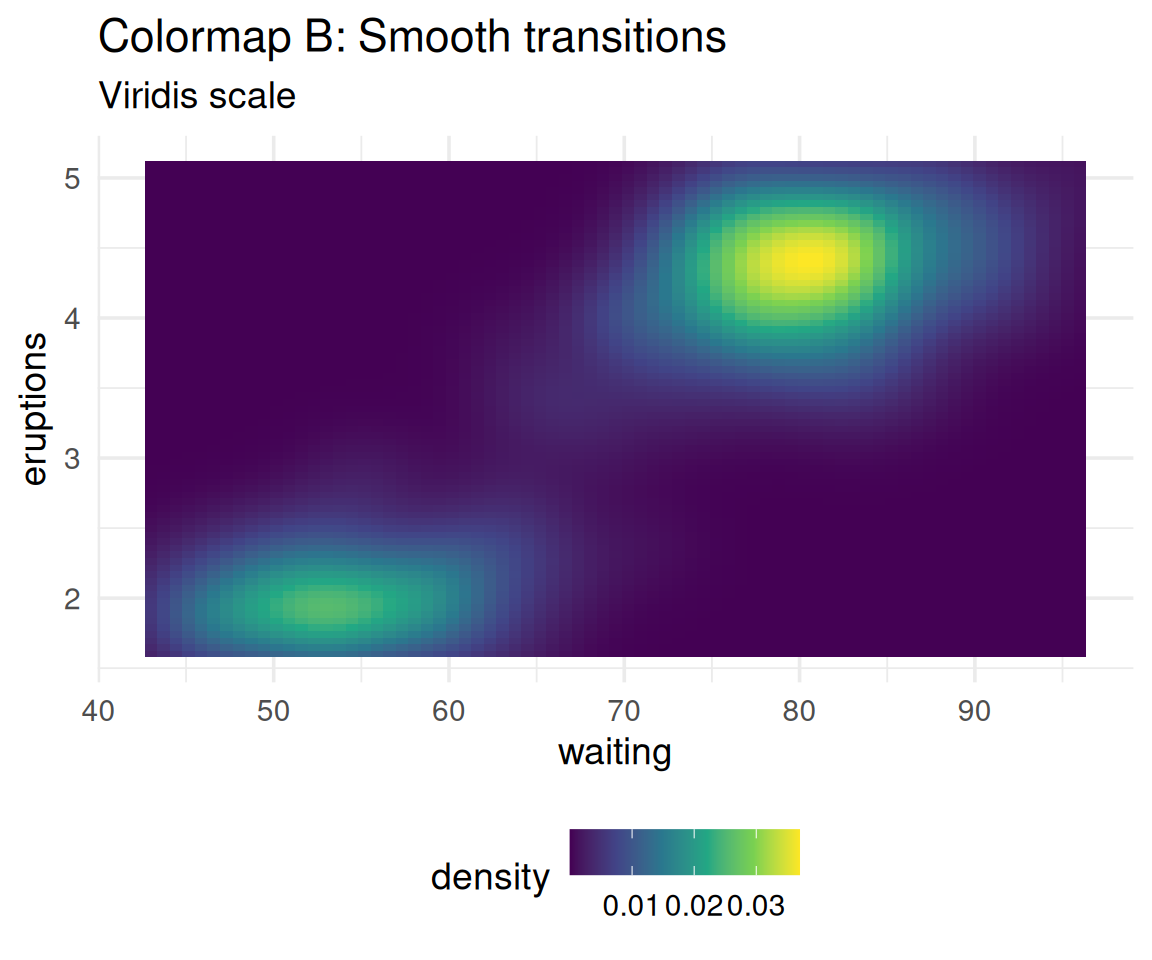
Which one shows the data more accurately?
The data is smooth, yet Colormap A creates false boundaries!
Perceptual Non-Uniformity: Demonstrated
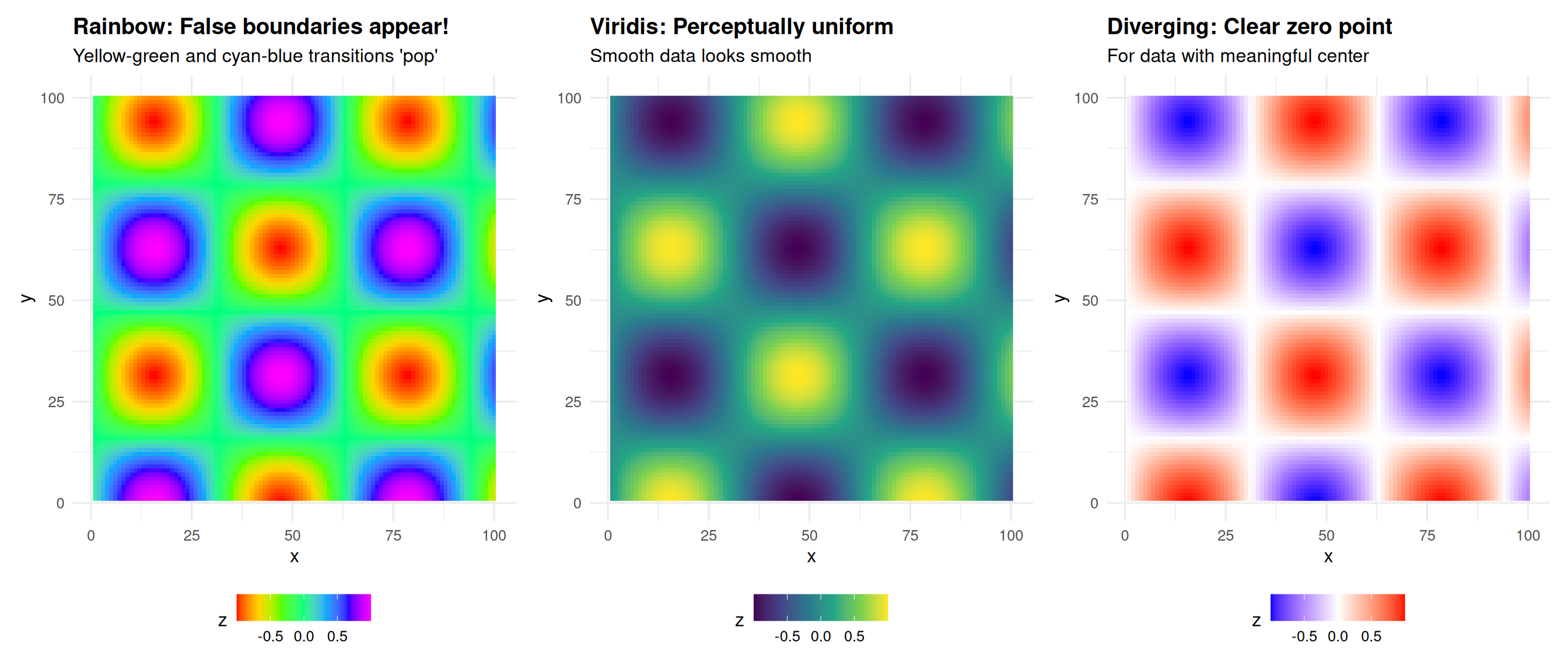
The data is perfectly smooth, yet rainbow creates artificial edges!
Real world consequences
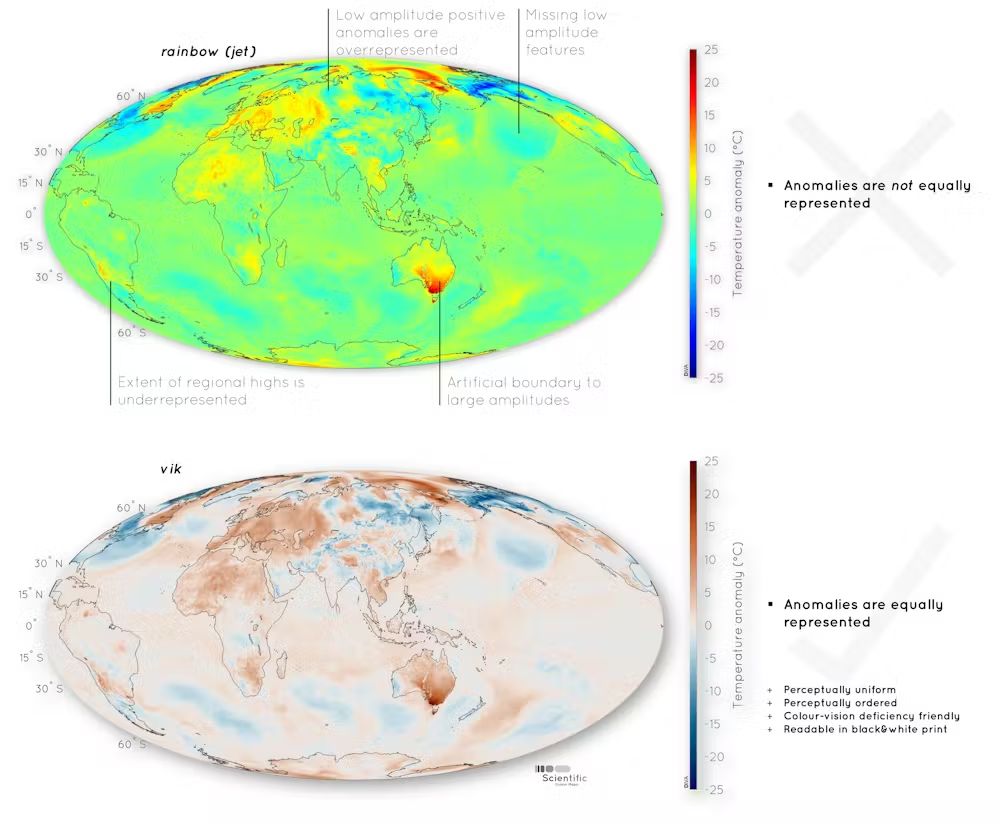
Not just spatial data
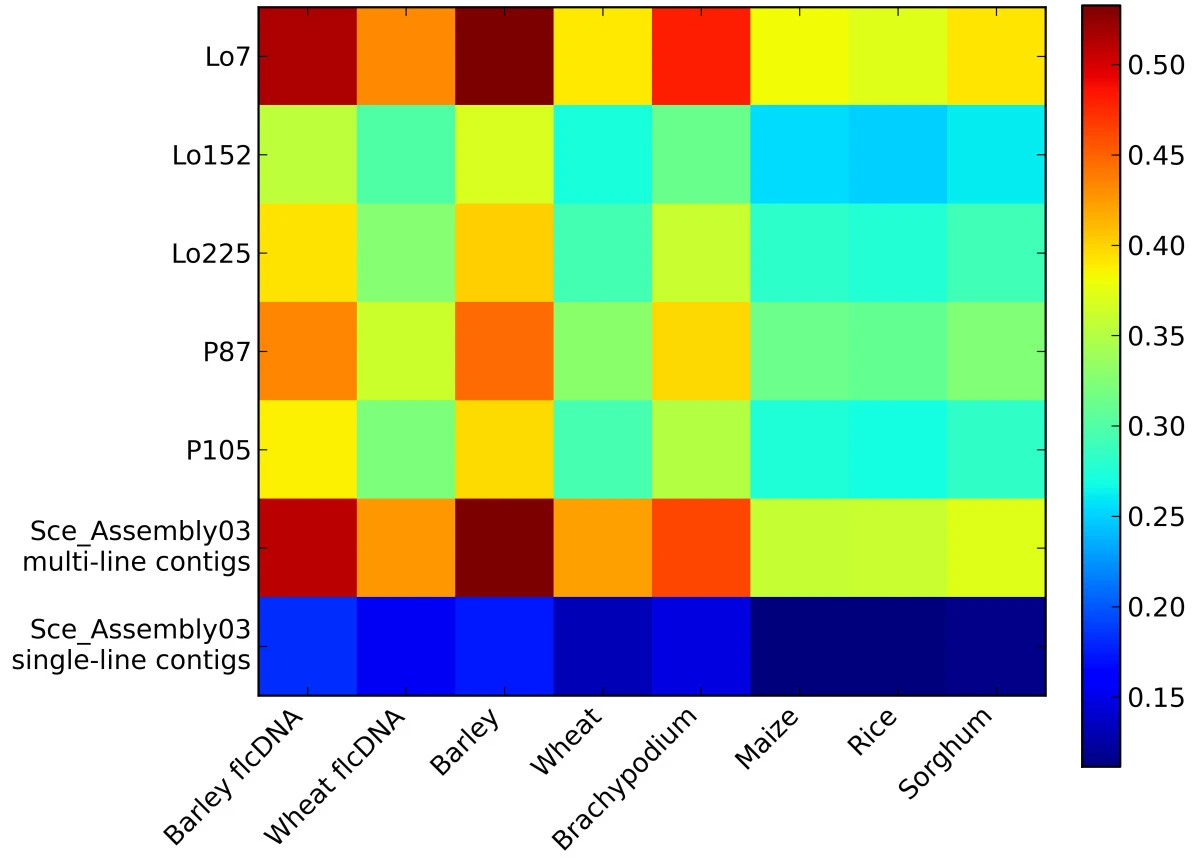

Figure from Haseneyer et al. (2011)
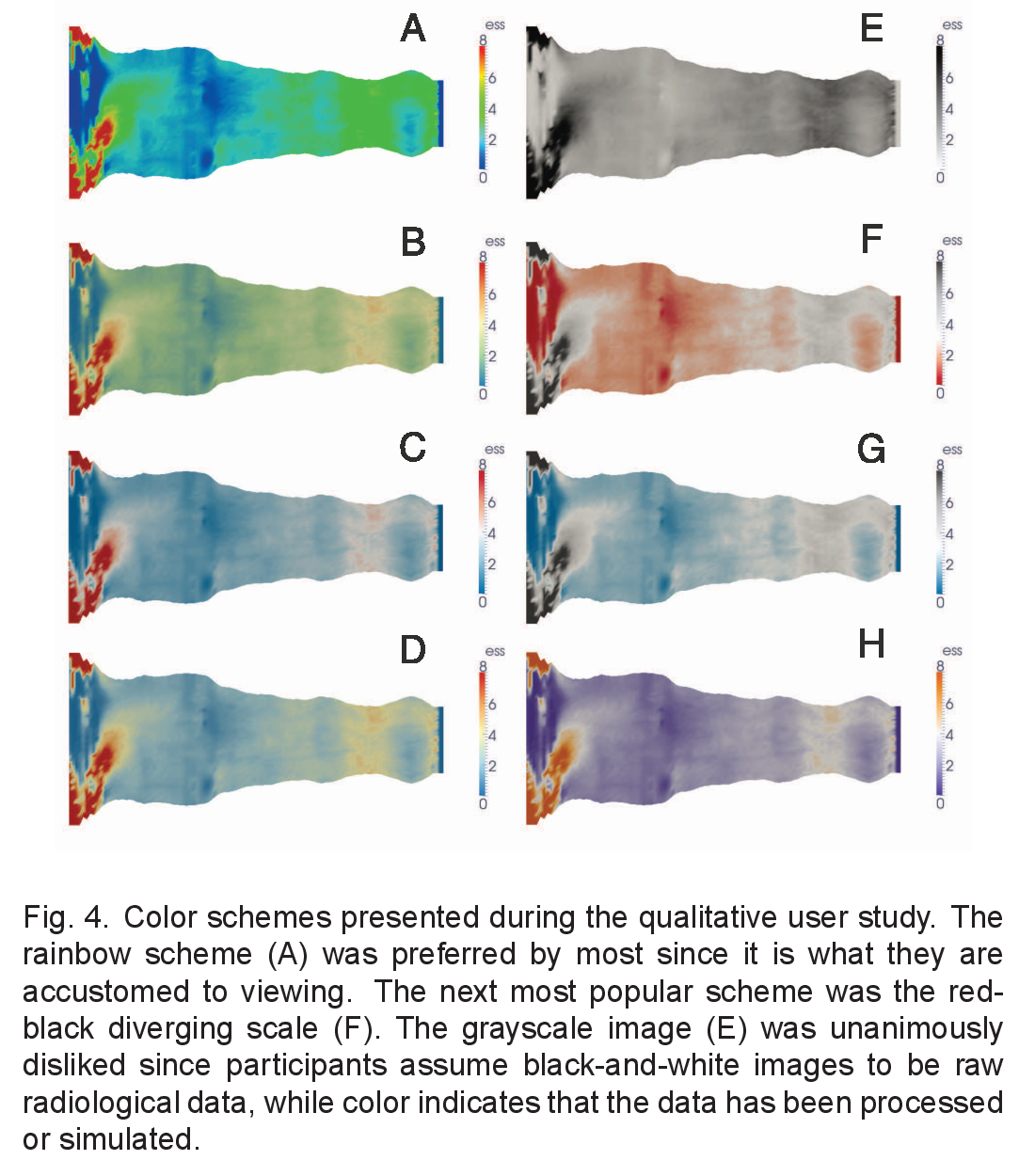
Medical Consequences
Lives at Stake
Borkin et al. (2011) - IEEE Visualization
Studied physicians diagnosing heart disease using medical imaging:
- Physicians using jet colormap: More errors, slower diagnosis
- Physicians using perceptually uniform colormaps: Fewer errors, faster
Why?
- Bright yellow appears more “intense” than dark red
- But dark red represents higher values (more critical condition)
- Perceptual bias leads to misdiagnosis
Reference: Borkin et al. (2011)
Comparison: Rainbow vs Better Alternatives
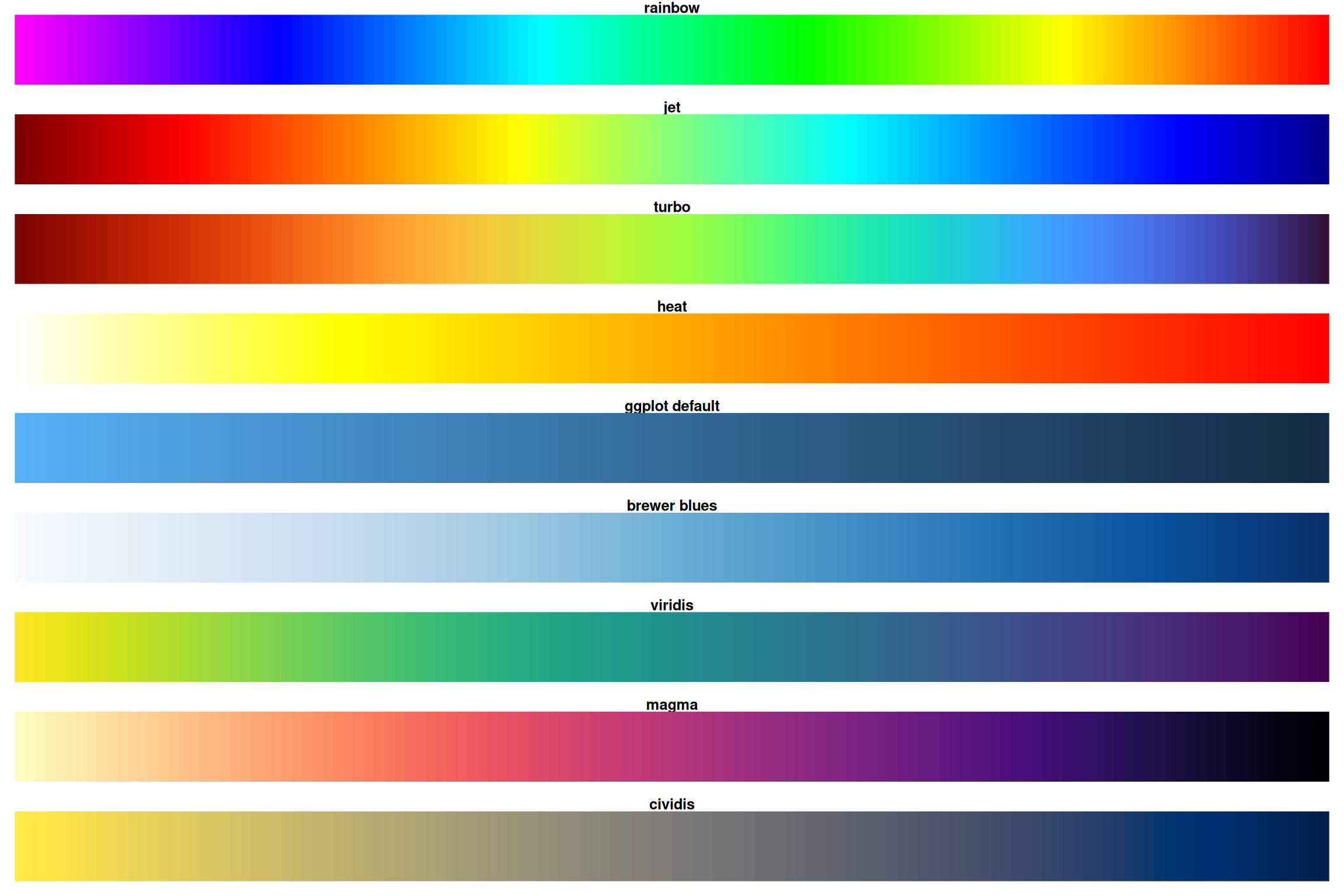

Notice how rainbow and heat have sharp transitions while viridis/magma are smooth!
Desaturated
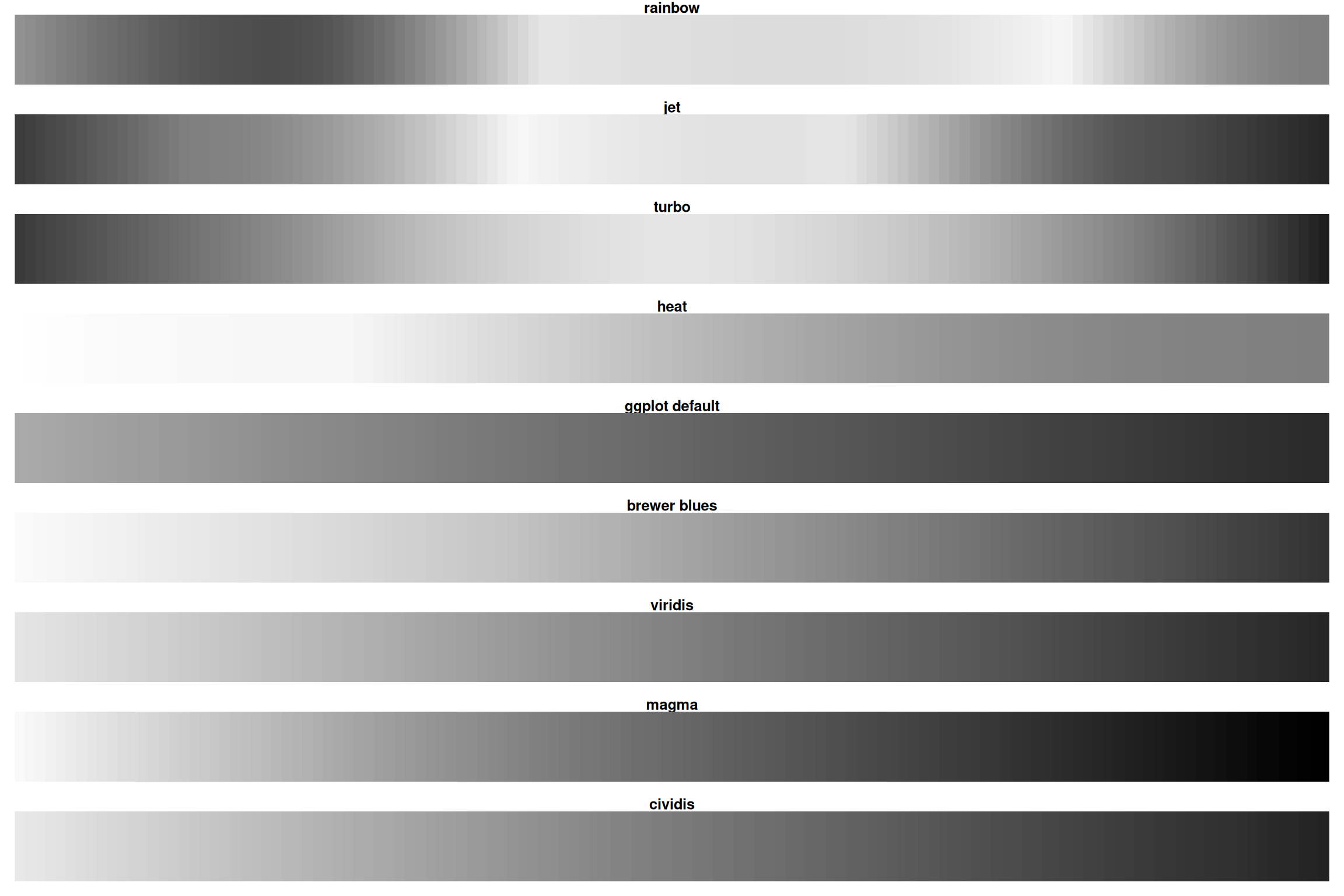

Rainbow loses all information when desaturated! Viridis/magma remain readable.
The Viridis Color Scales
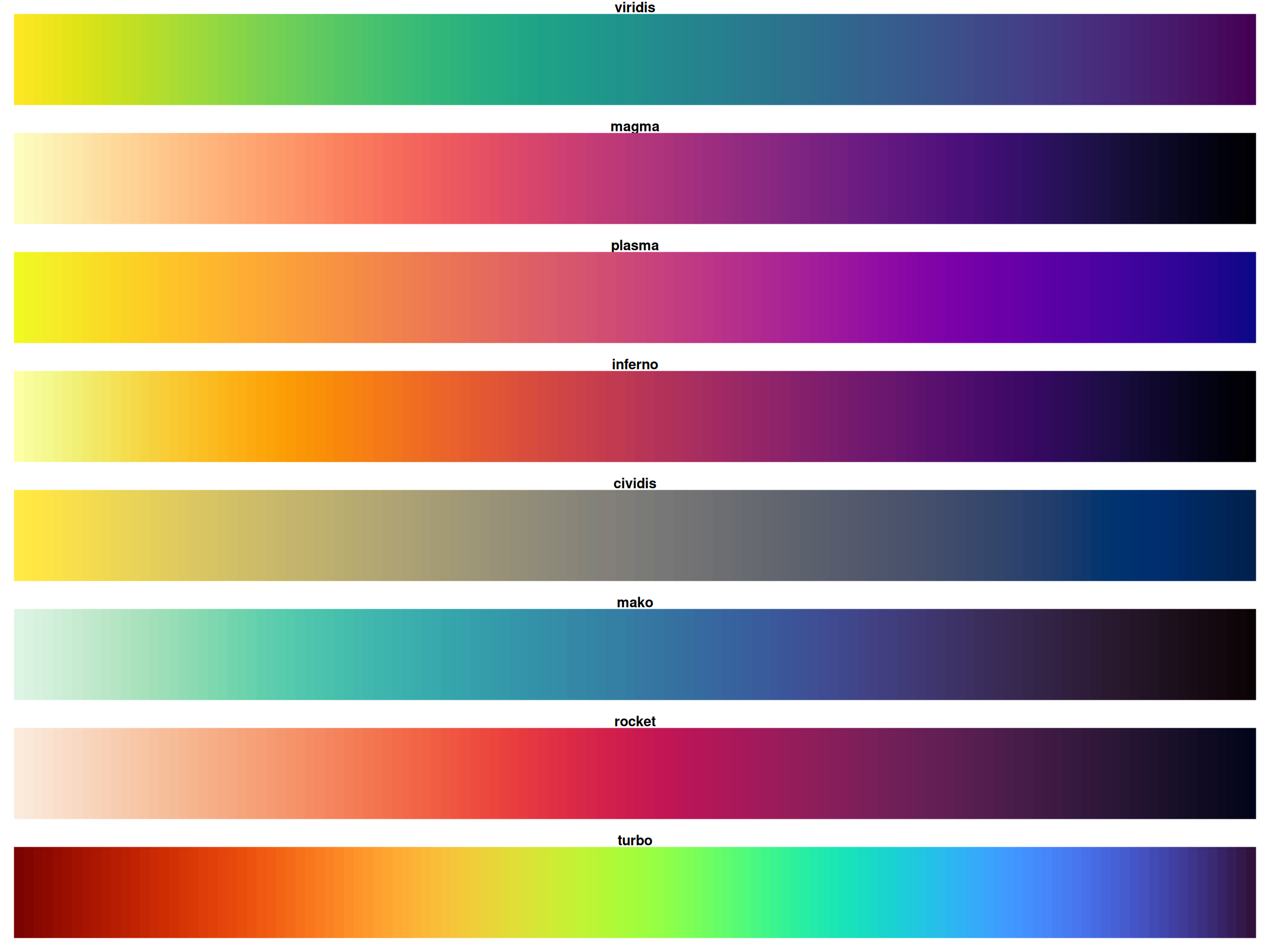

All viridis scales are perceptually uniform
The Viridis Color Scales, desaturated

Green-Blind Vision (Deuteranopia)
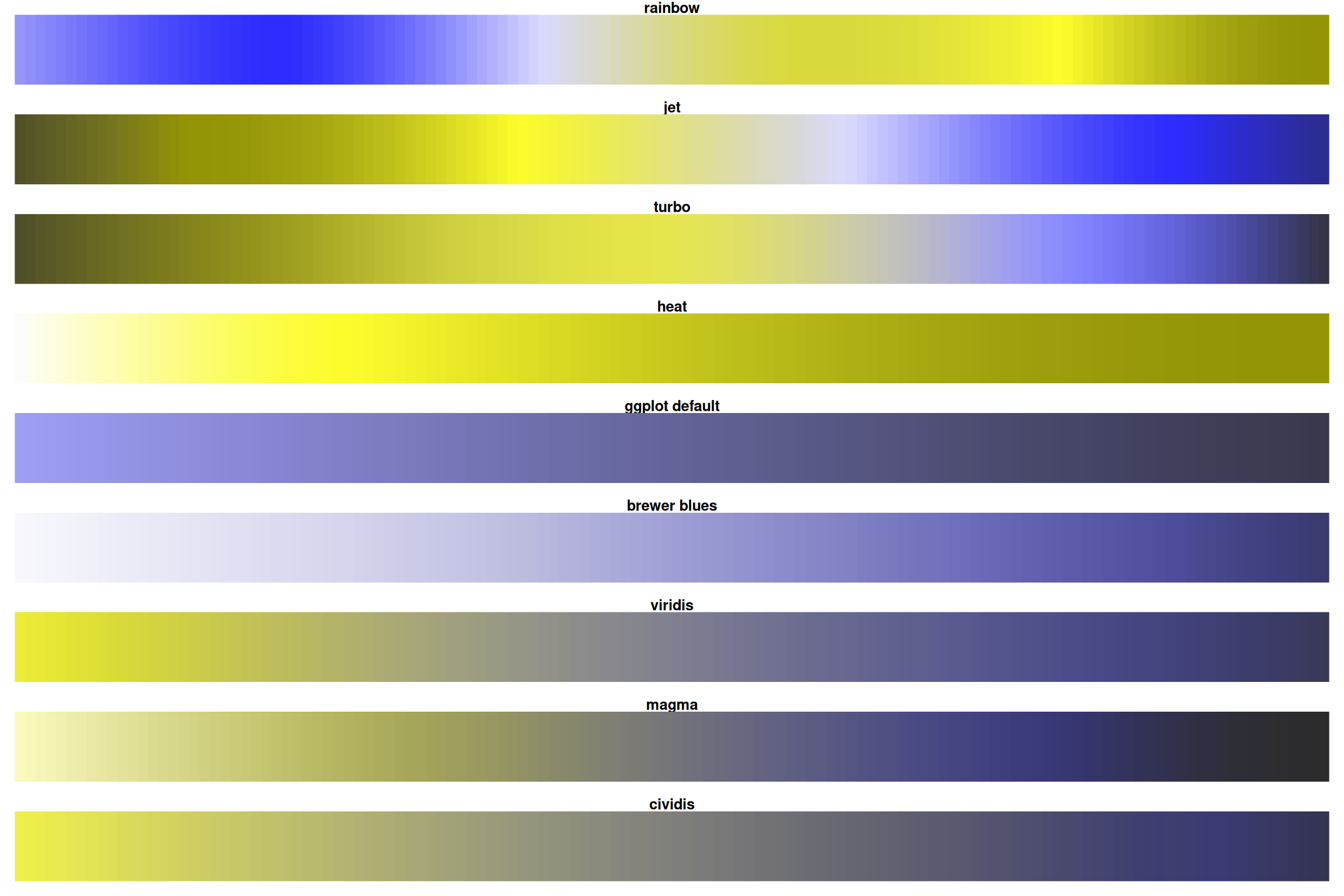

For ~8% of men, rainbow is nearly useless! Viridis/magma stay distinct.
Color Scale Comparison: Pros and Cons
|
Rainbow
|
Jet
|
Turbo
|
Heat
|
ggplot default
|
Brewer Blues
|
Viridis
|
Magma
|
Cividis
|
|
|---|---|---|---|---|---|---|---|---|---|
| Perceptually uniform | ❌ | ❌ | ✅ | ❌ | ✅ | ✅ | ✅ | ✅ | ✅ |
| Colorblind safe | ❌ | ❌ | ❌ | ❌ | ✅ | ✅ | ✅ | ✅ | ✅ |
| B&W/grayscale safe | ❌ | ❌ | ❌ | ❌ | ✅ | ✅ | ✅ | ✅ | ✅ |
| Good on projectors | ⚠️ | ⚠️ | ✅ | ⚠️ | ✅ | ✅ | ✅ | ✅ | ✅ |
| Print friendly | ❌ | ❌ | ✅ | ❌ | ✅ | ✅ | ✅ | ✅ | ✅ |
| Engaging colors | ✅ | ✅ | ✅ | ⚠️ | ⚠️ | ⚠️ | ✅ | ✅ | ⚠️ |
| Wide color range | ✅ | ✅ | ✅ | ⚠️ | ⚠️ | ⚠️ | ✅ | ✅ | ⚠️ |
| Recommendation | ❌ AVOID | ❌ AVOID | ⚠️ Careful | ❌ AVOID | OK | Good | ✅ DEFAULT | ✅ Great | ✅ Great |
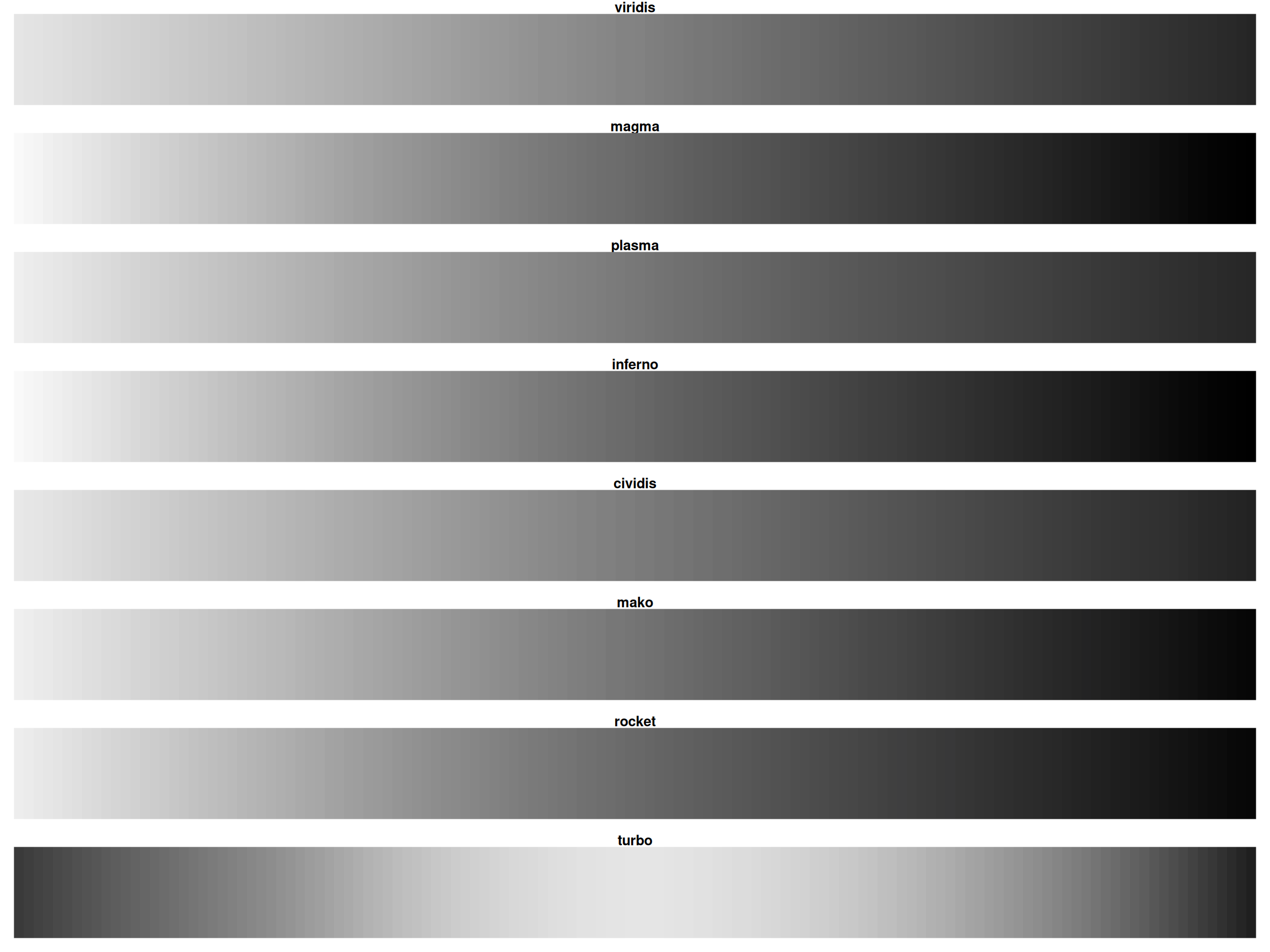
Viridis Family: Show All Options
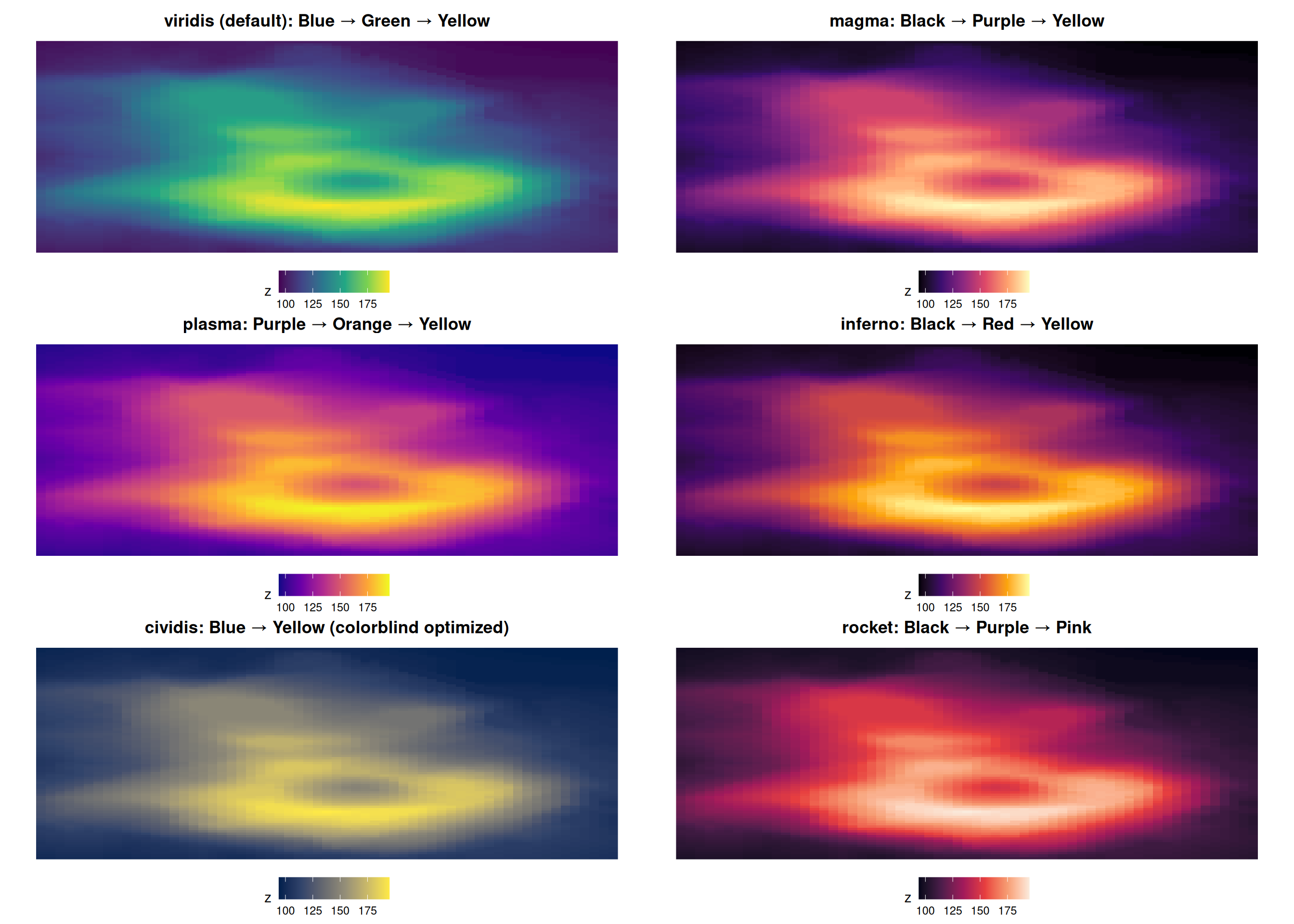
All are perceptually uniform and colorblind-friendly!
Recommendation for continous scales
For most use cases: Use Viridis (or Magma/Plasma variants)
When to use something else:
- Diverging data (has meaningful center): ColorBrewer diverging (RdBu, RdYlBu)
- High colorblind audience: Cividis
- Print-only publication: Brewer Blues or Greens
Never use: Rainbow or Jet
Colors for qualitative data
ColorBrewer: All Palettes
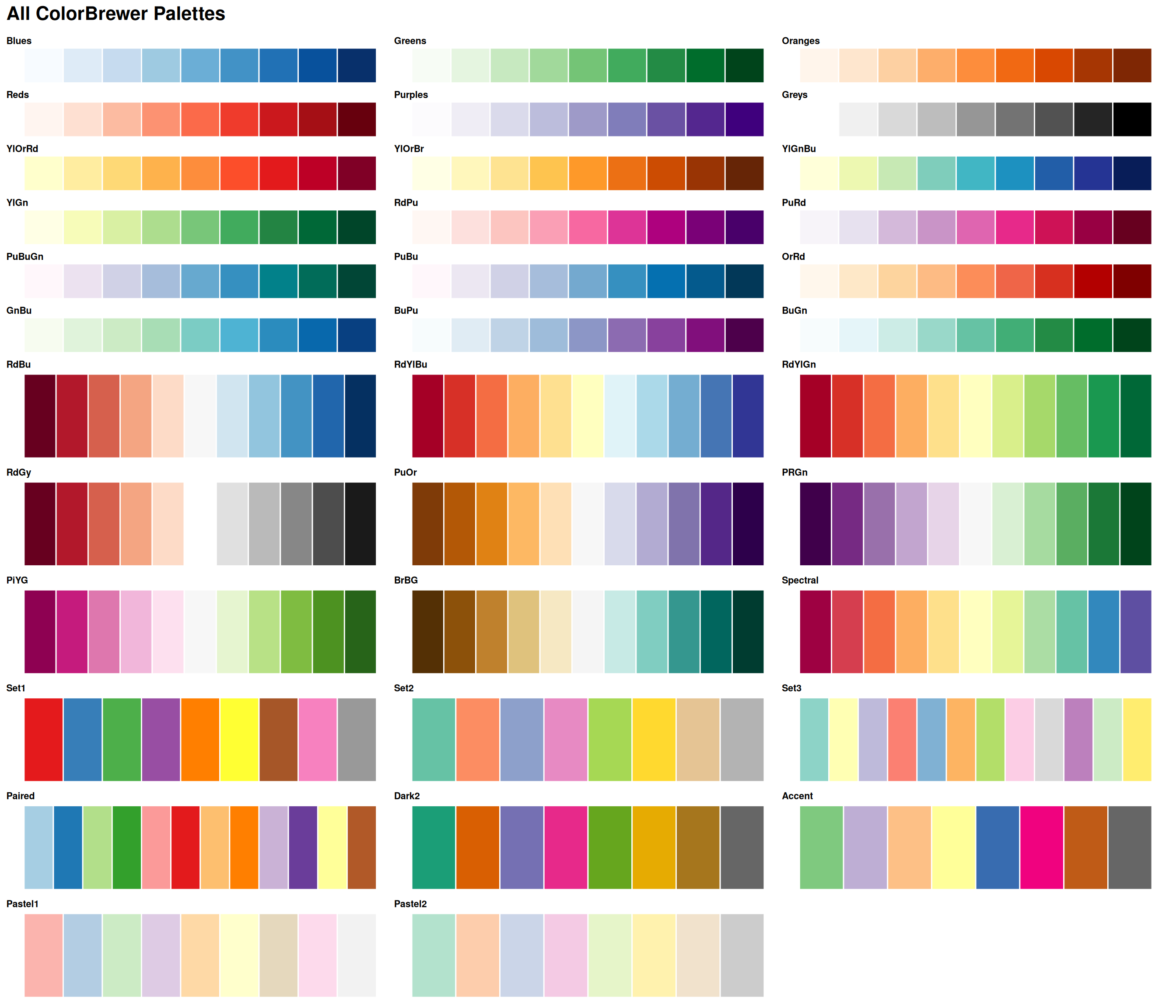

Explore all palettes at: colorbrewer2.org
ColorBrewer Website
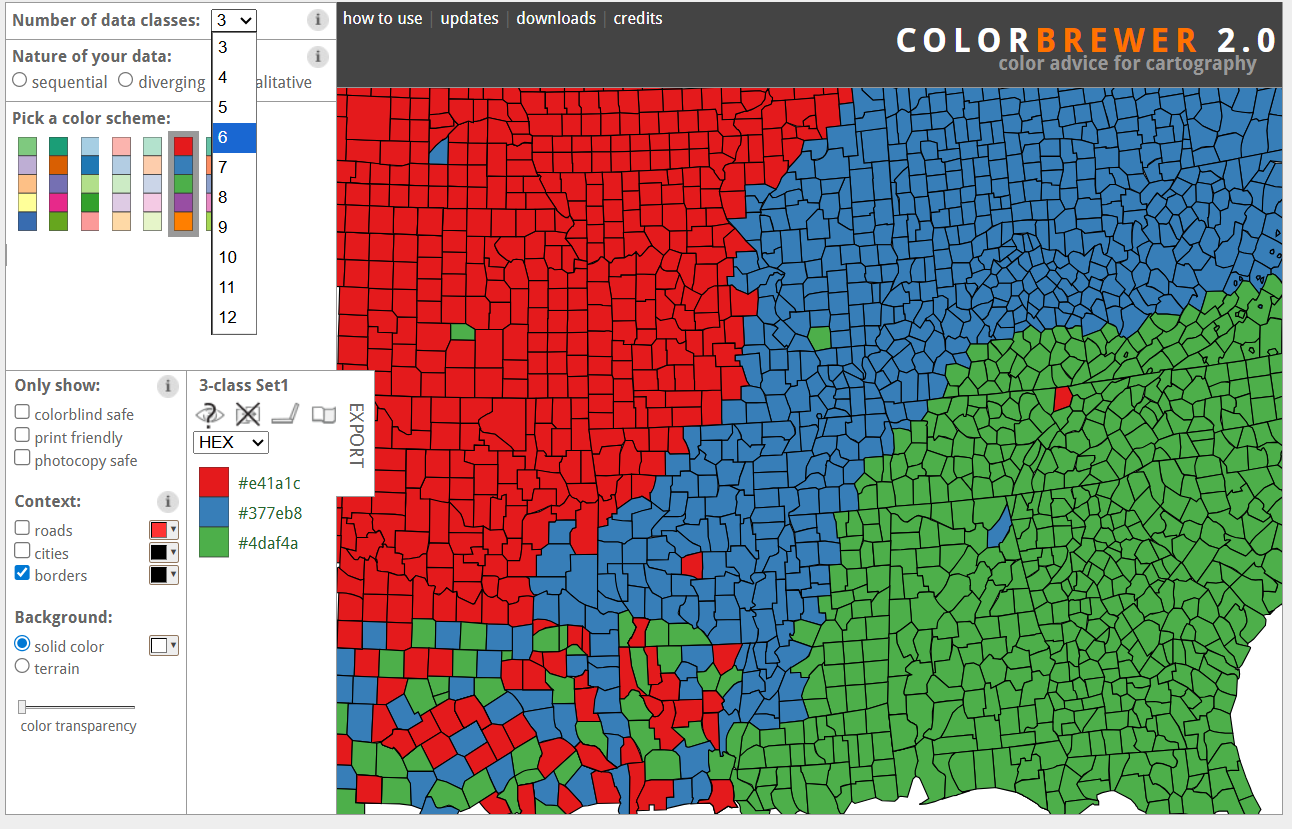
Yellow Color Warning
The Yellow Problem
Even though ColorBrewer includes yellow in some palettes (e.g., “YlOrRd”, “RdYlBu”, “Set1”):
Yellow has serious issues:
- Poor printing: Yellow can be nearly invisible on white paper
- Projection problems: On projected slides, yellow often washes out
- Low contrast: Yellow text on white background is unreadable
- Photocopying: Disappears when photocopied in B&W
Recommendation:
- For presentations: Avoid yellow-heavy palettes
- For print: Use darker yellows or oranges instead
- For text: NEVER use yellow text on light backgrounds
Removing yellow from Set1:
library(RColorBrewer)
# Set1 has yellow as the 6th color
set1_colors <- brewer.pal(9, "Set1")
set1_colors
# [1] "#E41A1C" "#377EB8" "#4DAF4A" "#984EA3" "#FF7F00" "#FFFF33" "#A65628" "#F781BF" "#999999"
# Remove yellow (position 6)
set1_no_yellow <- set1_colors[-6]
# Use in ggplot2
scale_color_manual(values = set1_no_yellow)Recommendations Summary
Best Practices for Color
- Continuous data: Viridis family
scale_fill_viridis_c()orscale_color_viridis_c()- Options: “viridis”, “magma”, “plasma”, “inferno”, “cividis”
- Diverging data (meaningful center): ColorBrewer
scale_fill_distiller(palette = "RdBu")for continuousscale_fill_gradient2()for custom diverging
- Categorical/qualitative data:
- Viridis:
scale_fill_viridis_d()orscale_color_viridis_d() - ColorBrewer:
scale_fill_brewer(palette = "Set2")(up to 8-12 categories)
- Viridis:
- AVOID:
- ❌ Rainbow/jet colormaps
- ❌ Red-green combinations (colorblind issue)
- ❌ Yellow text or yellow-heavy palettes (visibility issue)
- ALWAYS TEST:
- ✅ Grayscale conversion
- ✅ Colorblind simulation
- ✅ Print preview
Quick Reference: Code Examples
library(ggplot2)
# --- CONTINUOUS DATA ---
# Viridis continuous (best default)
ggplot(data, aes(x, y, fill = continuous_var)) +
geom_raster() +
scale_fill_viridis_c(option = "viridis") # or "magma", "plasma", "cividis"
# ColorBrewer sequential continuous
ggplot(data, aes(x, y, fill = continuous_var)) +
geom_raster() +
scale_fill_distiller(palette = "Blues") # or "YlOrRd", "Greens", etc.
# --- DIVERGING DATA (meaningful center) ---
# ColorBrewer diverging
ggplot(data, aes(x, y, fill = fold_change)) +
geom_raster() +
scale_fill_distiller(palette = "RdBu", direction = 1)
# Custom diverging
ggplot(data, aes(x, y, fill = fold_change)) +
geom_raster() +
scale_fill_gradient2(low = "blue", mid = "white", high = "red", midpoint = 0)
# --- CATEGORICAL/QUALITATIVE DATA ---
# Viridis discrete
ggplot(data, aes(x, y, color = category)) +
geom_point() +
scale_color_viridis_d(option = "viridis")
# ColorBrewer qualitative
ggplot(data, aes(x, y, color = category)) +
geom_point() +
scale_color_brewer(palette = "Set2") # or "Dark2", "Paired", etc.Heatmap Scaling
The Outlier Problem
Scenario: Metabolomics data (log2-transformed)
- Most metabolites: 6 to 10 (log2 scale)
- 1 outlier metabolite: ~10 (2^10 = 1000x higher!)
What happens with default scaling?
The extreme outlier compresses the color scale for all other values!
Visual Example: The Outlier Effect
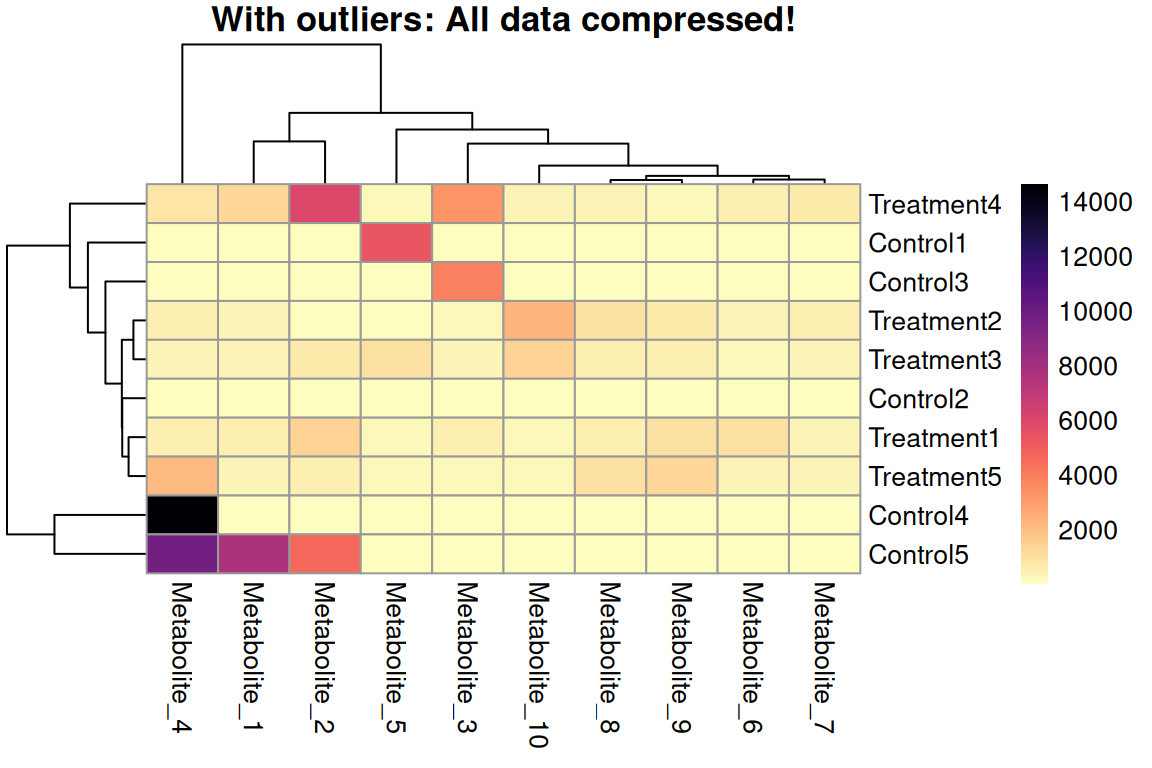
Without Outlier Handling
- Outliers dominate the color scale
- Most data compressed into narrow range
- Group differences invisible
- Patterns lost 😱
Solution 1: Robust MAD Scaling and Cutoffs
# Step 1: Robust scaling using MAD (Median Absolute Deviation)
expr_scaled <- expr_data %>%
mutate(across(-Sample, scale_mad))
# Step 2: Identify which values will be capped
expr_mat_scaled <- expr_scaled %>% column_to_rownames("Sample") %>% as.matrix()
capped_cells <- (expr_mat_scaled < -3) | (expr_mat_scaled > 3)
# Step 3: Cap at ±3 (meaningful after MAD scaling!)
expr_capped <- expr_scaled %>% mutate(across(-Sample, ~ pmin(pmax(.x, -3), 3)))
# Step 4: Create symmetric breaks centered at 0
max_abs <- max(abs(range(expr_capped[,-1])))
breaks <- seq(-max_abs, max_abs, length.out = 101)# Create asterisk markers for capped values (in original order)
asterisk_matrix <- matrix("", nrow = nrow(capped_cells), ncol = ncol(capped_cells))
asterisk_matrix[capped_cells] <- "*"
# Create final plot with asterisk markers
# display_numbers uses the original data order, clustering is applied automatically
p <- pheatmap(expr_capped %>% column_to_rownames("Sample"),
main = "MAD-scaled + capped at ±3 (* = capped)",
color = colorRampPalette(rev(brewer.pal(11, "RdBu")))(100),
breaks = breaks, scale = "none",
display_numbers = asterisk_matrix,
number_color = "black",
fontsize_number = 14,
silent = TRUE)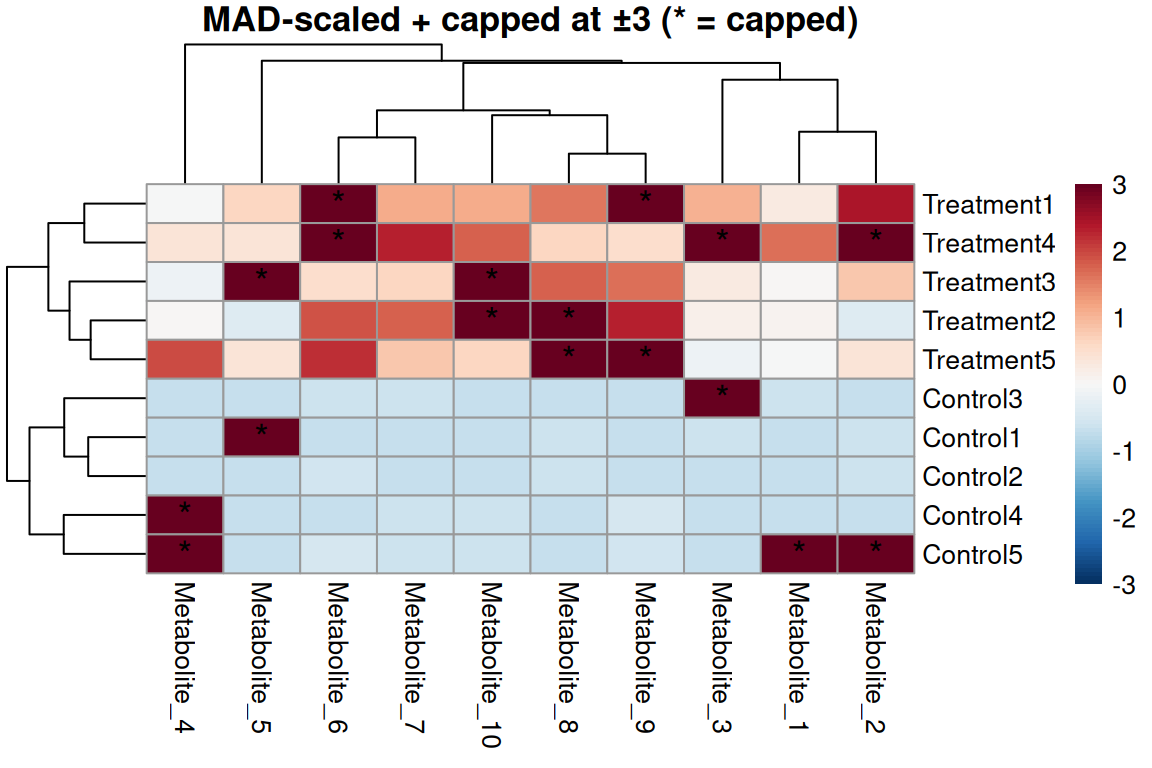
Robust scaling approach
- Use MAD instead of SD - not affected by outliers
- Center by median (robust)
- Scale by MAD (Median Absolute Deviation)
- Then cap at ±3 MAD (~99% of normal data)
- Outliers now exceed threshold!
- Set
scale = "none"(already scaled!)
Solution 2: Range scaling and Quantile-Based cut-off
# Step 1: Identify values outside 5-95 percentiles PER COLUMN
expr_mat_raw <- expr_data %>% column_to_rownames("Sample") %>% as.matrix()
capped_cells_q <- apply(expr_mat_raw, 2, function(x) {
q_lower <- quantile(x, 0.05, na.rm = TRUE)
q_upper <- quantile(x, 0.95, na.rm = TRUE)
(x < q_lower) | (x > q_upper)
})
# Step 2: Cap at 5th and 95th percentiles PER COLUMN (metabolite)
expr_capped <- expr_data %>%
mutate(across(-Sample, ~ cap_quantiles(.x, lower = 0.05, upper = 0.95)))
# Step 3: Range scaling (min-max normalization to [0,1])
expr_quantile <- expr_capped %>% mutate(across(-Sample, ~ (.x - min(.x)) / (max(.x) - min(.x))))# Create asterisk markers for capped values (in original order)
asterisk_matrix_q <- matrix("", nrow = nrow(capped_cells_q), ncol = ncol(capped_cells_q))
asterisk_matrix_q[capped_cells_q] <- "*"
# Create final plot with asterisk markers
p <- pheatmap(expr_quantile %>% column_to_rownames("Sample"),
main = "Capped at 5-95 percentiles + range-scaled (* = capped)",
color = rev(viridis::magma(100)),
display_numbers = asterisk_matrix_q,
number_color = "white",
fontsize_number = 14,
silent = TRUE)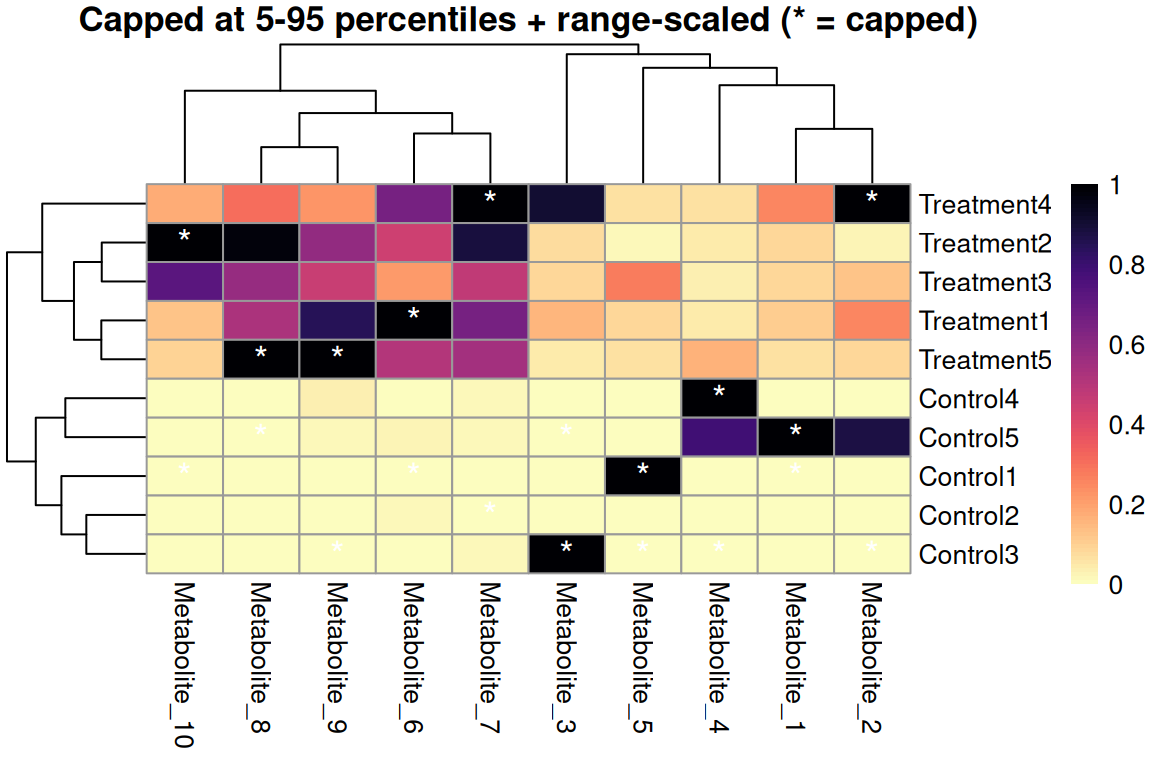
Quantile capping + range scaling
- Cap first to remove outliers per metabolite
- Then range scale to use full [0,1] color scale
- More robust to outliers than variance scaling
- Good for non-normal data
- Common quantiles: 5-95% or 2-98%
Solution 3: Log Transformation
For positive values only (e.g., counts, intensities)
# Log transform BEFORE plotting (metabolomics data is positive-only)
expr_log_sol4 <- expr_data %>%
mutate(across(-Sample, ~ log2(.x + 1))) # +1 to handle zeros
p <- pheatmap(expr_log_sol4 %>% column_to_rownames("Sample"),
main = "Log2 transformed metabolite intensities",
color = rev(viridis::magma(100)),
silent = TRUE)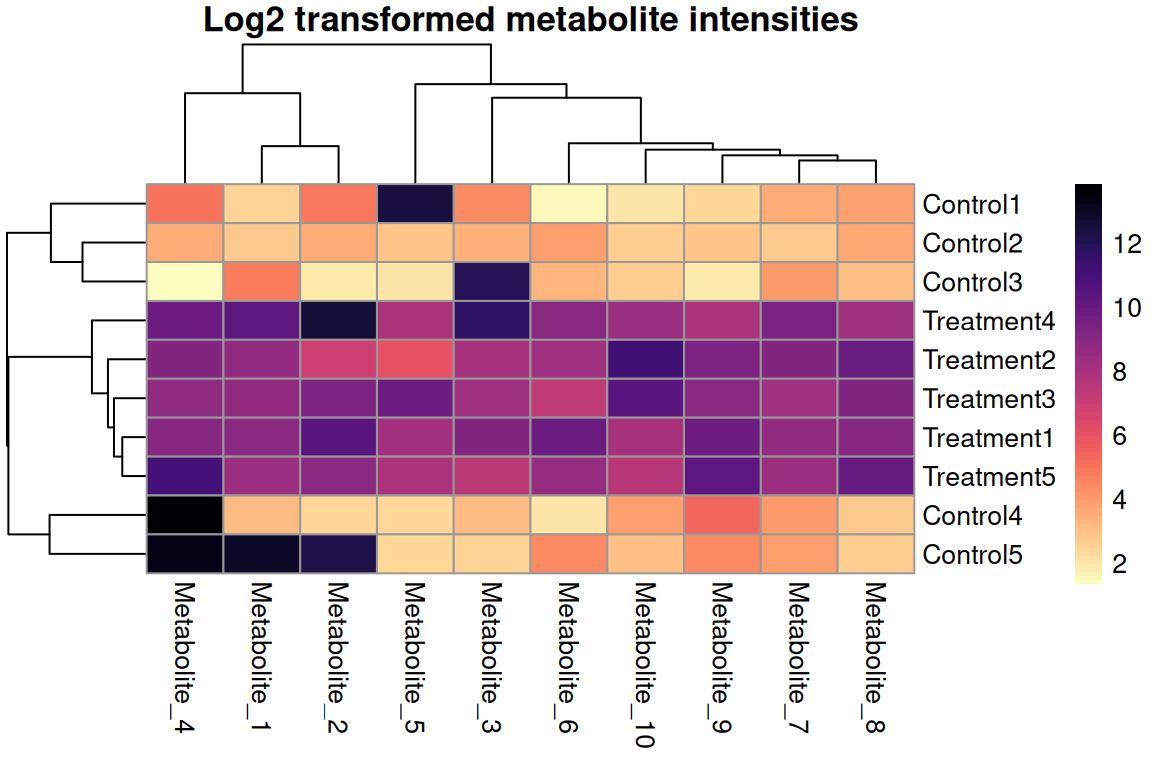
For count/intensity data
- Compresses wide ranges
- Add +1 to handle zeros
- Common for RNA-seq, proteomics
- Use log2, log10, or ln
The Dendrogram Scaling Trap
Hidden Technical Issue
Critical R bug: Functions like heatmap(), heatmap.2(), and heatplot() have a dangerous inconsistency:
- The
scaleparameter affects color visualization - But NOT dendrogram calculation!
Result: Dendrograms cluster on unscaled data while colors show scaled data!
P.S: pheatmap() seems to apply scaling and cropping before clustering!
Why Scaling Matters for Clustering
The Problem: High-variance features dominate correlations
Without scaling, features with large values dominate sample correlations!
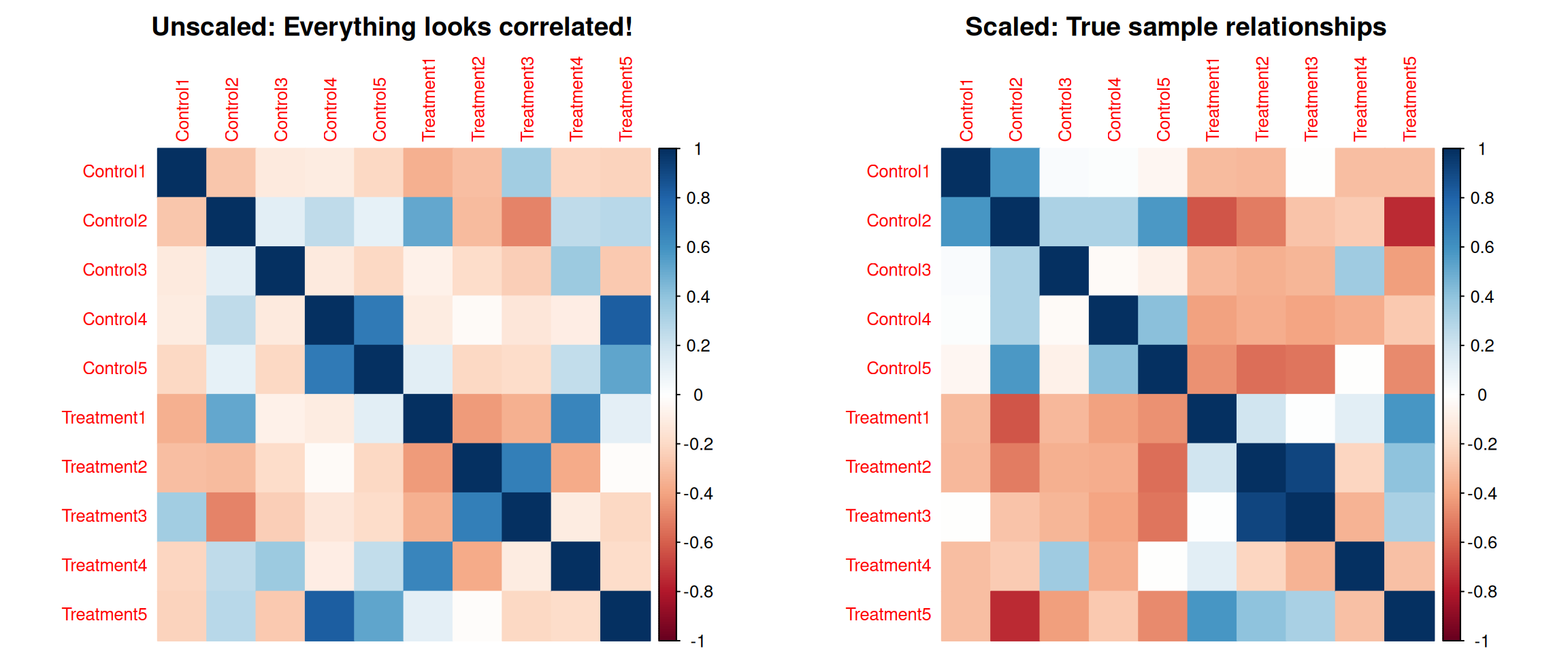
Why Scaling Matters for Clustering (2)
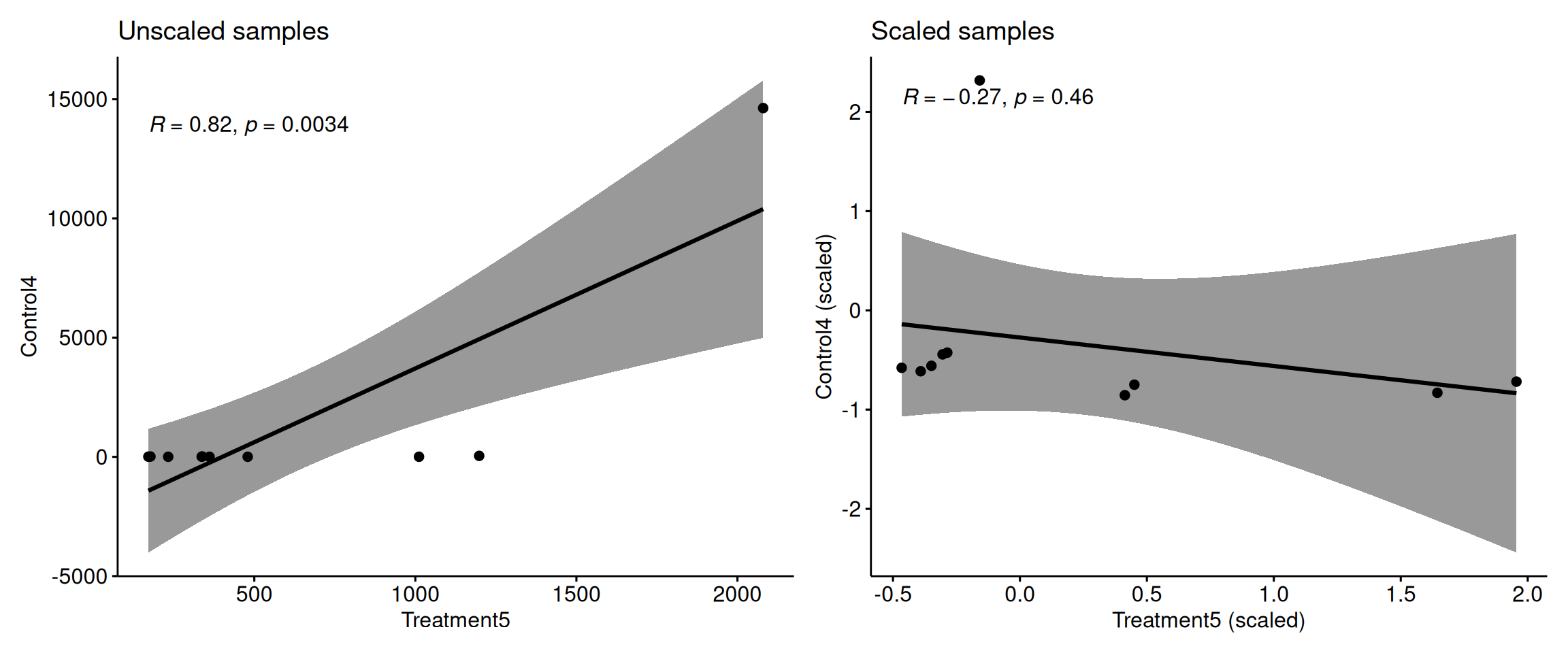
Key point: Without scaling, high-variance metabolites completely dominate the correlation calculation between samples!
Scaling ensures all metabolites contribute equally to sample clustering.
Using massageR::heat.clust
Better Approach: heat.clust
The massageR package provides heat.clust() which handles scaling and dendrogram calculation correctly in one step!
Key advantages:
- Scales data and calculates dendrograms together
- Controls exactly where limits are applied (data and/or dendrograms)
- Returns pre-computed dendrograms
- Works seamlessly with pheatmap
heat.clust with pheatmap
library(massageR)
# Convert tibble to matrix for heat.clust
expr_matrix <- expr_data %>% column_to_rownames("Sample") %>% as.matrix()
# Use heat.clust with robust MAD scaling
z <- heat.clust(expr_matrix,
scaledim = "column", # Scale by column
zlim = c(-3, 3), # Cap at ±3 MAD
zlim_select = c("dend", "outdata"), # Apply to both
reorder = c(), # Reorder dendrograms off for consistency
distfun = function(x) dist(x),
hclustfun = function(x) hclust(x, method = "complete"),
scalefun = scale_mad) # Use MAD scaling instead of default
max_abs <- max(abs(range(z$data)))
breaks <- seq(-max_abs, max_abs, length.out = 101)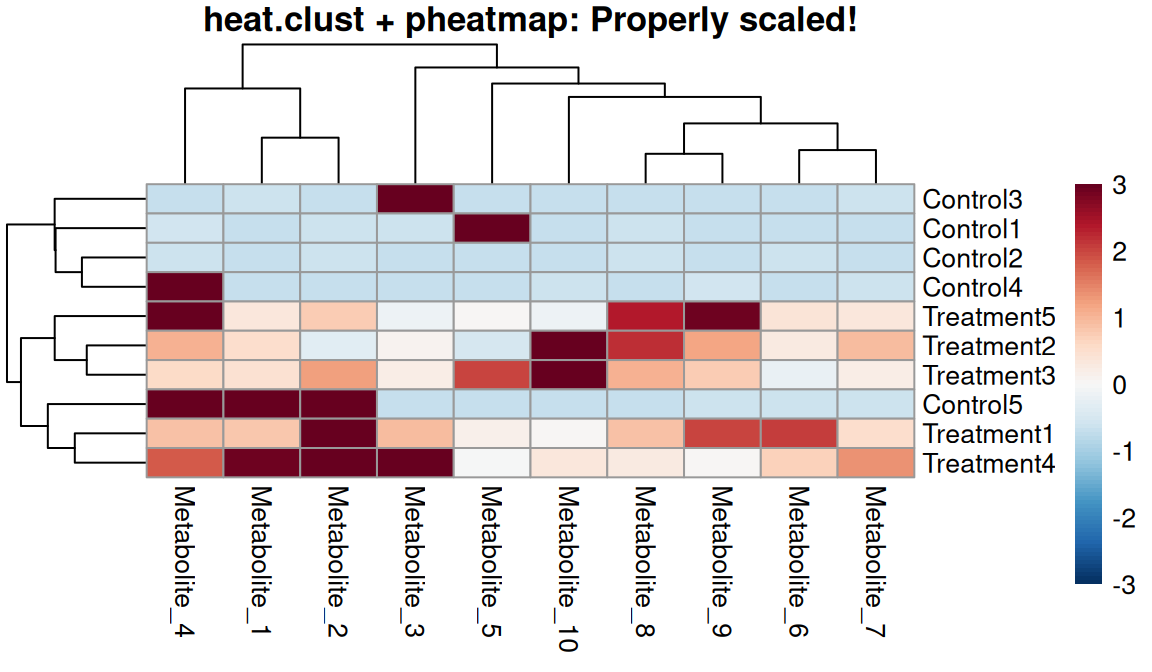
One-step workflow
- Scales data automatically
- Calculates dendrograms on scaled data
- Caps at specified zlim
- Returns everything needed for pheatmap
- Ensures consistency throughout
Comparison: Before and After
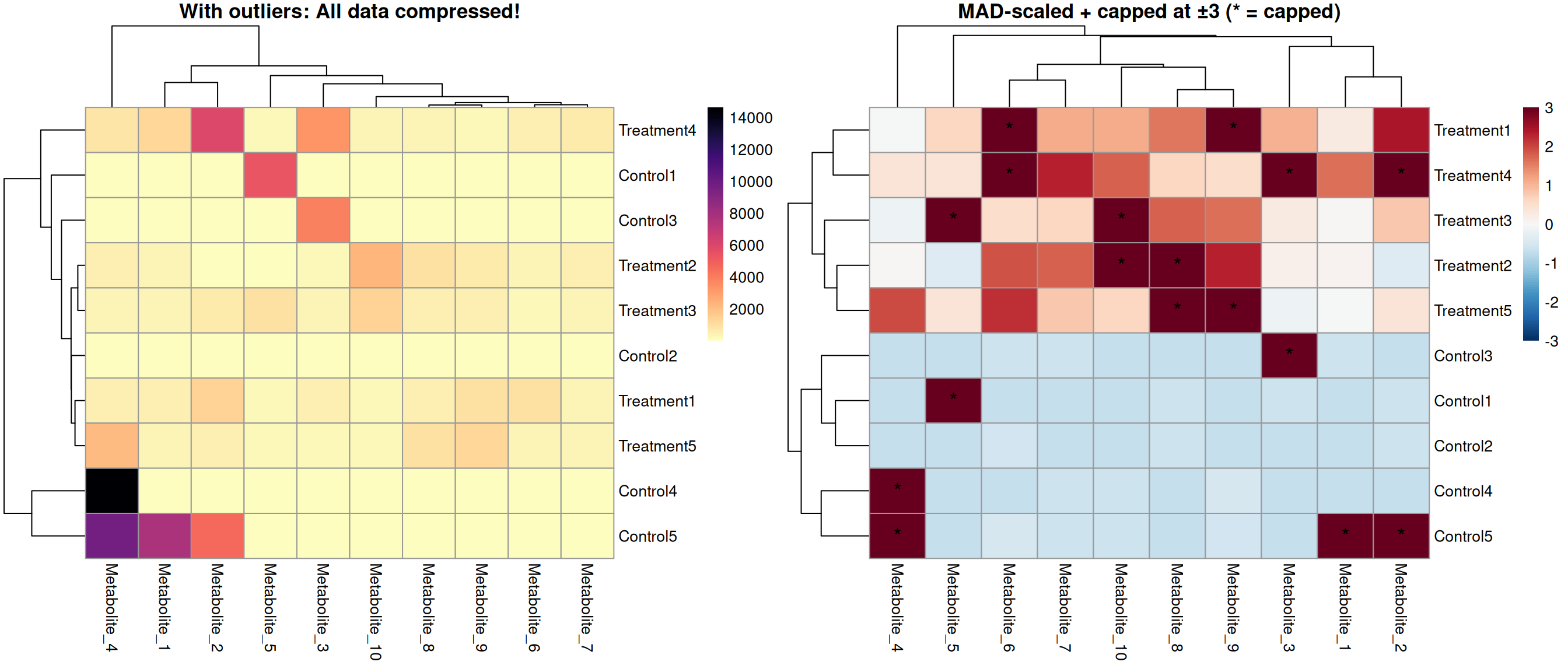
Best Practices & Recommendations
Workflow
- Inspect data distribution before making heatmap
- Consider log transformation
- Scale data BEFORE passing to heatmap function
- Use MAD (robust) if data has outliers:
(x - median(x)) / mad(x) - Use SD if data is clean:
scale(x)
- Use MAD (robust) if data has outliers:
- Cap extremes
- Cap at ±3 MAD for robust scaling with outliers
- Cap using quantiles (5-95%) if using range scaling
- Calculate dendrograms on the same scaled and capped data
- Consider
massageR::heat.clust()for automatic proper scaling workflow - Use appropriate palette:
- Often centering data highlights contrasts
- Diverging color scale for data that has been centered (red-white-blue)
- Sequential color scale for one-directional data (viridis, magma)
When NOT to Cap
- If outliers are biologically meaningful (rare events)
- Small datasets where each value matters
- When you want to highlight extreme values
Which image format for which purpose?
What is a Raster Image?
Raster graphics are grids of colored pixels
- Stores individual pixel colors: RGB(255, 128, 64)
- Fixed resolution measured in DPI (Dots Per Inch)
- More pixels = higher resolution = larger file size
- Cannot be scaled up without quality loss
What is a Vector Image?
Vector graphics use mathematical descriptions
- Mathematical formulas define shapes and lines
- “Draw a line from point (0,0) to (10,10)”
- “Create a circle with center (5,5) and radius 3”
- Infinitely scalable without quality loss
Infinite Resolution
Because vectors are mathematical formulas, they can be scaled to any size without losing quality. The curve is defined by equations, not pixels!
Vector vs. Raster Comparison
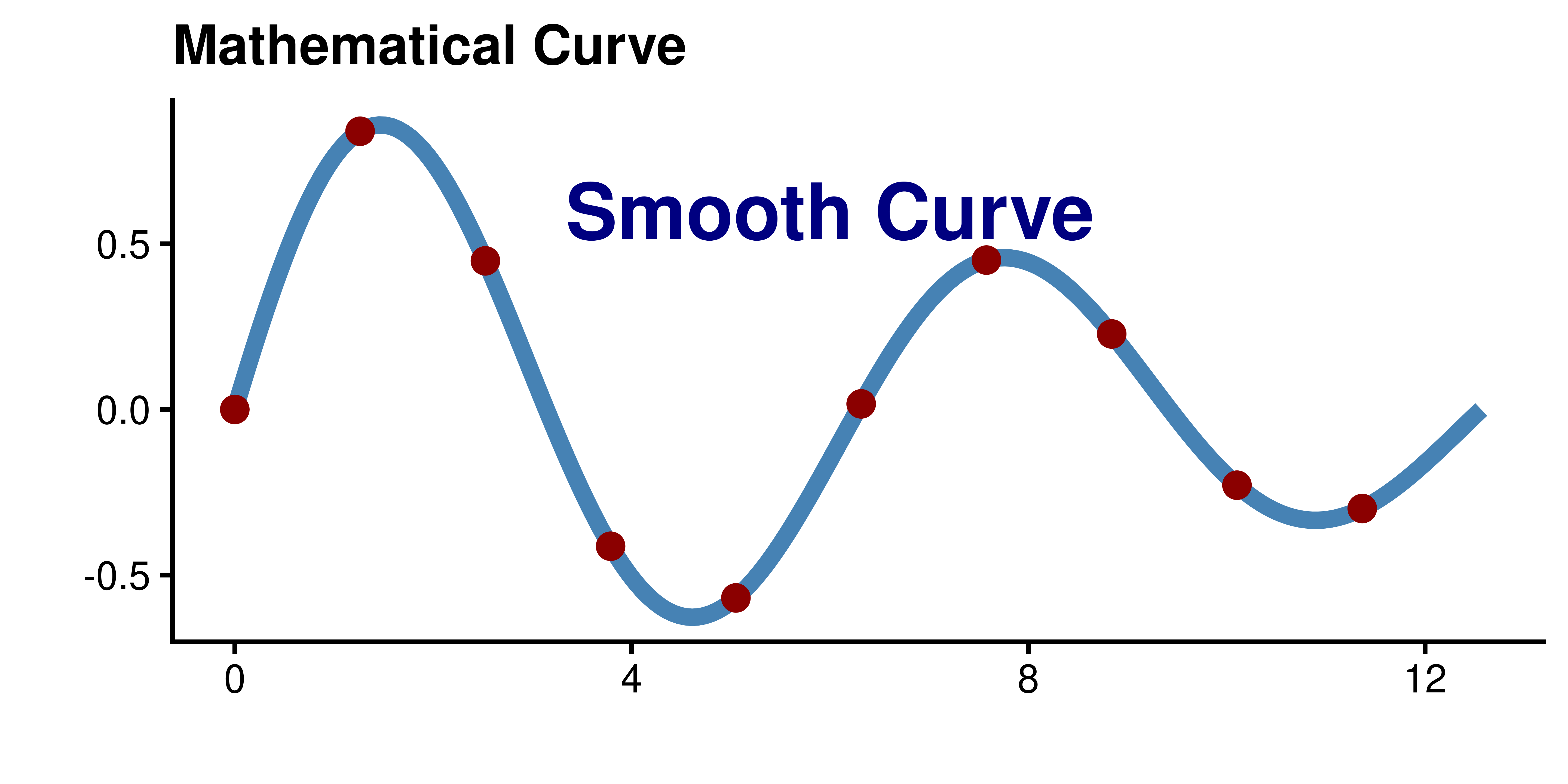
Vector vs. Raster Comparison (2)

| Aspect | Vector (PDF, SVG, EPS) | Raster (PNG, TIFF, JPG) |
|---|---|---|
| Definition | Mathematical formulas | Grid of pixels |
| Scalability | Infinite resolution | Fixed resolution (DPI) |
| File Size | Small (formulas compact) | Large (all pixels stored) |
| Best For | Plots, diagrams, text, screenshots of websites | Photos, screenshots |
| Editability | Easy to edit paths | Pixel-level editing only |
| Text Quality | Always crisp | Can become blurry |
Container Formats: PDF and TIFF
PDF and TIFF are Containers!
Both PDF and TIFF can contain EITHER vector OR raster data:
- PDF: Can contain vector graphics, raster images, or both
- TIFF: Usually raster, but can embed vector data
TIFF Compression Options:
| Type | Description | Use Case |
|---|---|---|
| Uncompressed | No compression (huge files) | Archival |
| LZW | Lossless compression | Publications |
| ZIP | Lossless compression | Publications |
| JPEG | Lossy compression | Web (avoid for science) |
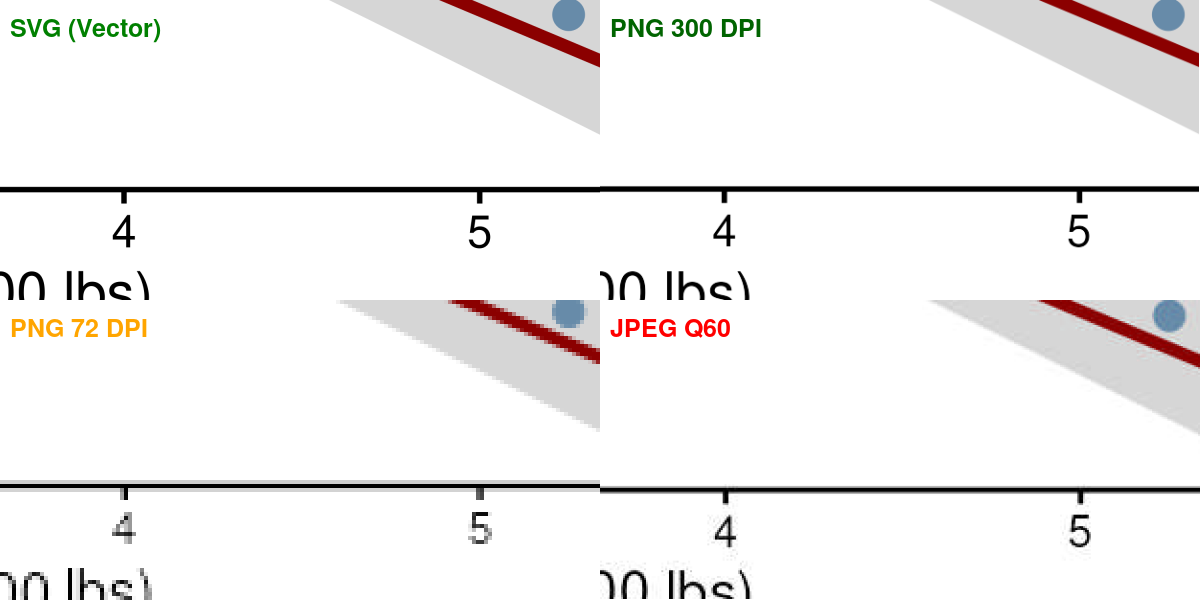
JPEG Compression Artifacts

Format Properties Comparison
| Format | Type | Compression | Container? | Notes |
|---|---|---|---|---|
| Vector + Raster | Lossless | Yes | Can embed both vector and raster data | |
| EPS | Vector | Lossless | Yes | Older format required by some journals. Use device = "eps" in ggsave(). PDF is preferred when accepted. |
| SVG | Vector | Lossless | No | XML-based, web-native |
| PNG | Raster | Lossless | No | Supports transparency |
| TIFF | Raster | Lossless or Lossy | Yes | Multiple pages, various compression options |
| JPEG | Raster | Lossy | No | Best for photos only |
| WebP | Raster | Lossless or Lossy | No | Modern web format, smaller than PNG/JPEG |
Container Formats
Container formats can hold multiple types of data or multiple images:
- PDF: Can mix vector graphics, raster images, fonts, and text
- EPS: Can embed fonts, preview images, and vector data
- TIFF: Can contain multiple pages/images with different compression
Non-container formats store a single image with one encoding type.
File Format Comparison
Real world horror story
This is the graphical abstract
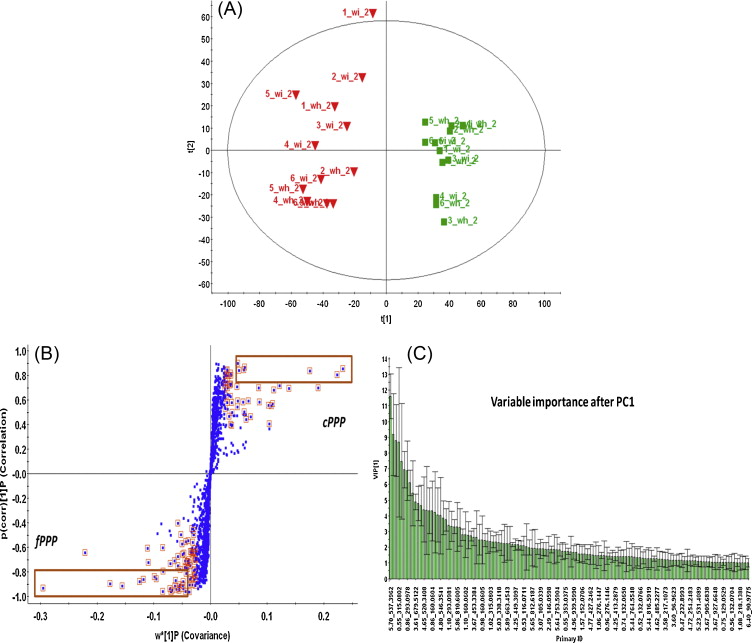
This was one of the figures
Vector Screenshots from Websites
Capturing Website Content as Vector Graphics
Print to PDF as you see the website
- Open the webpage in Google Chrome
- Open Chrome DevTools (
F12) - Click “⋮”
- Click “More Tools”
- click “Rendering”
- Set “Emulate CSS media type” to “screen”
- “Print”to PDF
Why This Matters
- Text stays crisp - Perfect for including web-based figures
- Smaller file size - Vector PDF is compact
- Editable - Can extract or edit elements in PDF editor
- Publication quality - No pixelation when zoomed
Examples:
- Document online data visualization examples
- Include web-based tools in presentations and papers
Format Selection: Decision Tree
%%{init: {'theme':'dark', 'themeVariables': {'edgeLabelBackground':'#1a1a1a', 'primaryTextColor':'#fff', 'secondaryTextColor':'#fff', 'tertiaryTextColor':'#fff'}}}%%
flowchart LR
A[What type of image?] --> B{Photo}
A --> C{Screenshot}
A --> D{Generated figure/<br/>website snapshot}
B --> B1{Where will it<br/>be used?}
B1 -->|Publication| B2[TIFF LZW<br/>300+ DPI]
B1 -->|Presentation/Web| B3[JPEG/WebP <br/> 150+ DPI]
C --> C1{Where will it<br/>be used?}
C1 -->|Publication| C2[TIFF LZW<br/>300+ DPI]
C1 -->|Presentation/Web| C3[PNG<br/>150 DPI]
D --> D1{Publication or<br/>presentation?}
D1 -->|Publication| D2{Vector support?}
D1 -->|Presentation/Web| D3[SVG or PNG]
D2 -->|Yes| D4[PDF / SVG / EPS<br/>All equivalent vectors]
D2 -->|No| D5[TIFF 600+ DPI LZW<br/>PNG not supported]
style A fill:#5dade2,color:#fff
style B fill:#5dade2,color:#fff
style C fill:#5dade2,color:#fff
style D fill:#5dade2,color:#fff
style B1 fill:#5dade2,color:#fff
style C1 fill:#5dade2,color:#fff
style D1 fill:#5dade2,color:#fff
style D2 fill:#5dade2,color:#fff
style D4 fill:#2ecc71,color:#fff
style D5 fill:#5dade2,color:#fff
style D3 fill:#9b59b6,color:#fff
style B2 fill:#3498db,color:#fff
style B3 fill:#e74c3c,color:#fff
style C2 fill:#3498db,color:#fff
style C3 fill:#9b59b6,color:#fff
Last notes
- Make sure that any art you include is vector based!
- Composing your figures in Powerpoint is OK - but use high quality source-material
- BioRender material is typically vector based
Text and Element Sizing
The Key Insight
The Smart Way to Control Text Size
Don’t manually set font sizes for every element!
Instead, use smaller figure dimensions to make text appear larger relative to the plot.
Then use vector formats (SVG/PDF) for infinite resolution.
Visual Demonstration
base_size Effect (theme_classic(base_size = x))
base_size = 8
base_size = 11 (default)
base_size = 14
Other elements scale relative to base_size:
axis.title:1.1× base_sizeaxis.text:0.8× base_sizelegend.text:0.8× base_size
Use base_size as the PRIMARY adjustment - only customize individual elements if needed
The Right Workflow
Three-Step Process
- Set dimensions to match final output size
- Journal single column: ~3.5 inches (Always check specific journal guidelines!)
- Journal double column: ~7 inches
- Presentation: ~10 inches
- Adjust
base_sizeif needed- Only fine-tune if text still too large/small
- Adjust individual elements if needed
- Use
theme()to customize specific text sizes - Only if steps 1-2 don’t achieve desired result
- Use
Why This Works
- Text/points/lines have fixed sizes
- If the canvas is small they appear larger
When you must adjust some sizes individually
Vector vs Raster: Key Differences
Vector Formats (SVG, PDF)
Dimensions control text/element proportions, not quality!
- Scale infinitely without quality loss
- DPI is ignored
- What matters: aspect ratio & relative proportions
Raster Formats (PNG, TIFF)
DPI controls pixel count and quality!
- Fixed resolution, can pixelate when scaled
- DPI matters (300+ for print)
- pixels = width × DPI
ggplot2 Themes
The Default Problem
# Default ggplot2 theme
p <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE) +
labs(title = "Default theme_gray()",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon",
color = "Cylinders") +
theme(plot.title = element_text(face = "bold"))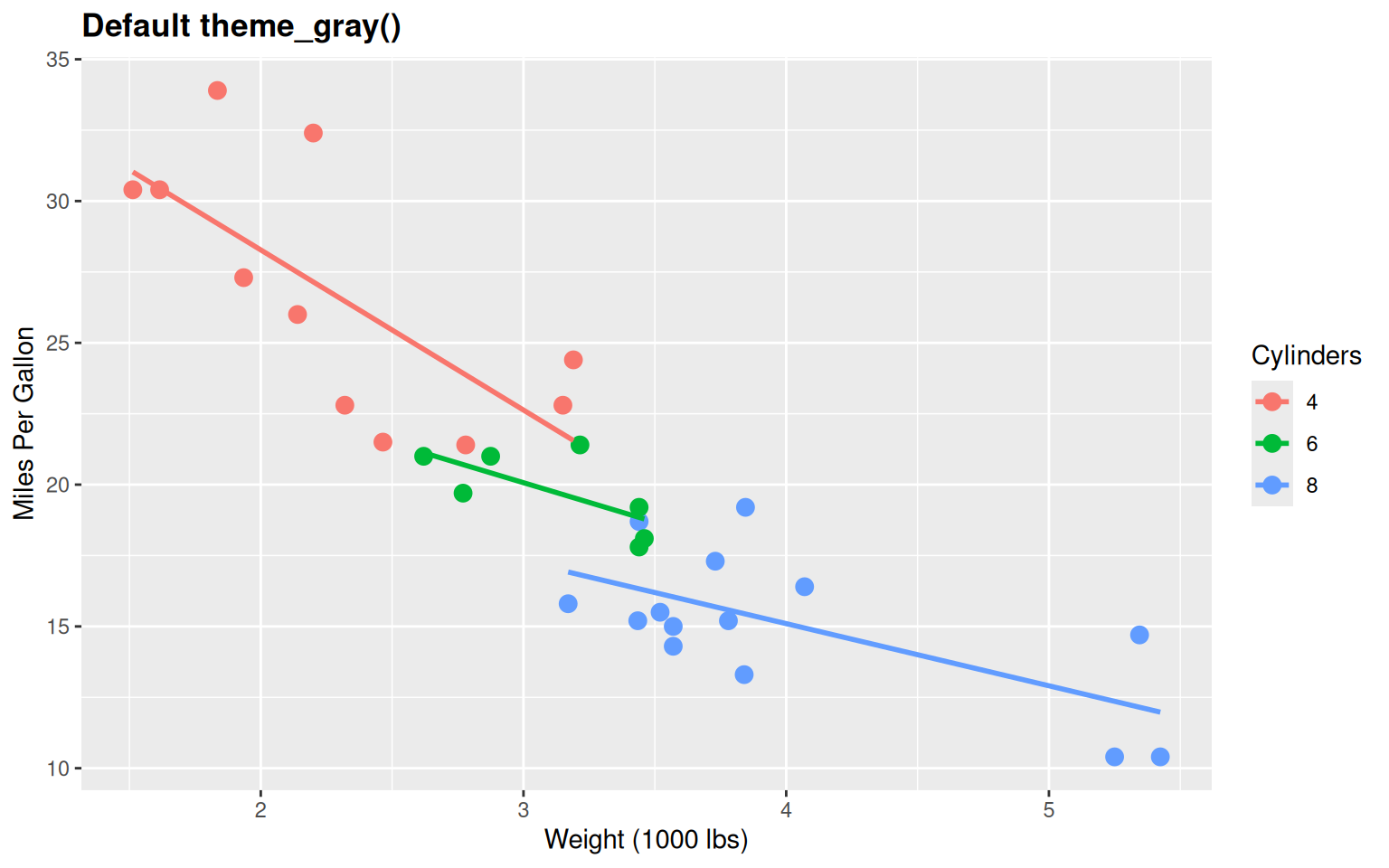
Problems:
- Gray background (wastes ink, unprofessional)
- Too much “chart junk”
- Not publication-ready
Built-in Theme: theme_bw()
p <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE) +
labs(title = "theme_bw() - White background, black border",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon",
color = "Cylinders") +
theme_bw() +
theme(plot.title = element_text(face = "bold"))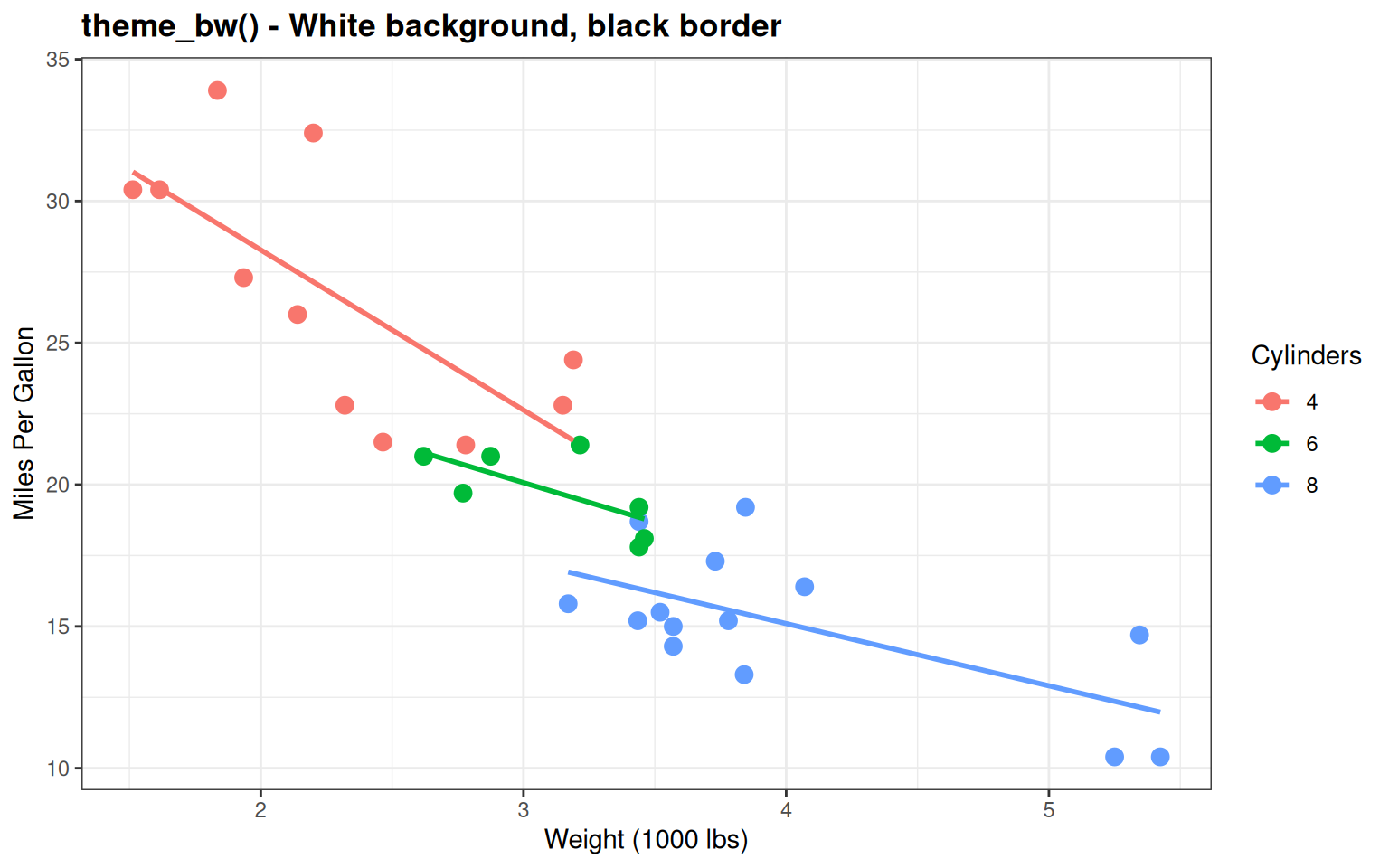
Good for publications - Clean with reference gridlines
Built-in Theme: theme_classic()
p <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE) +
labs(title = "theme_classic() - No gridlines, clean axes",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon",
color = "Cylinders") +
theme_classic() +
theme(plot.title = element_text(face = "bold"))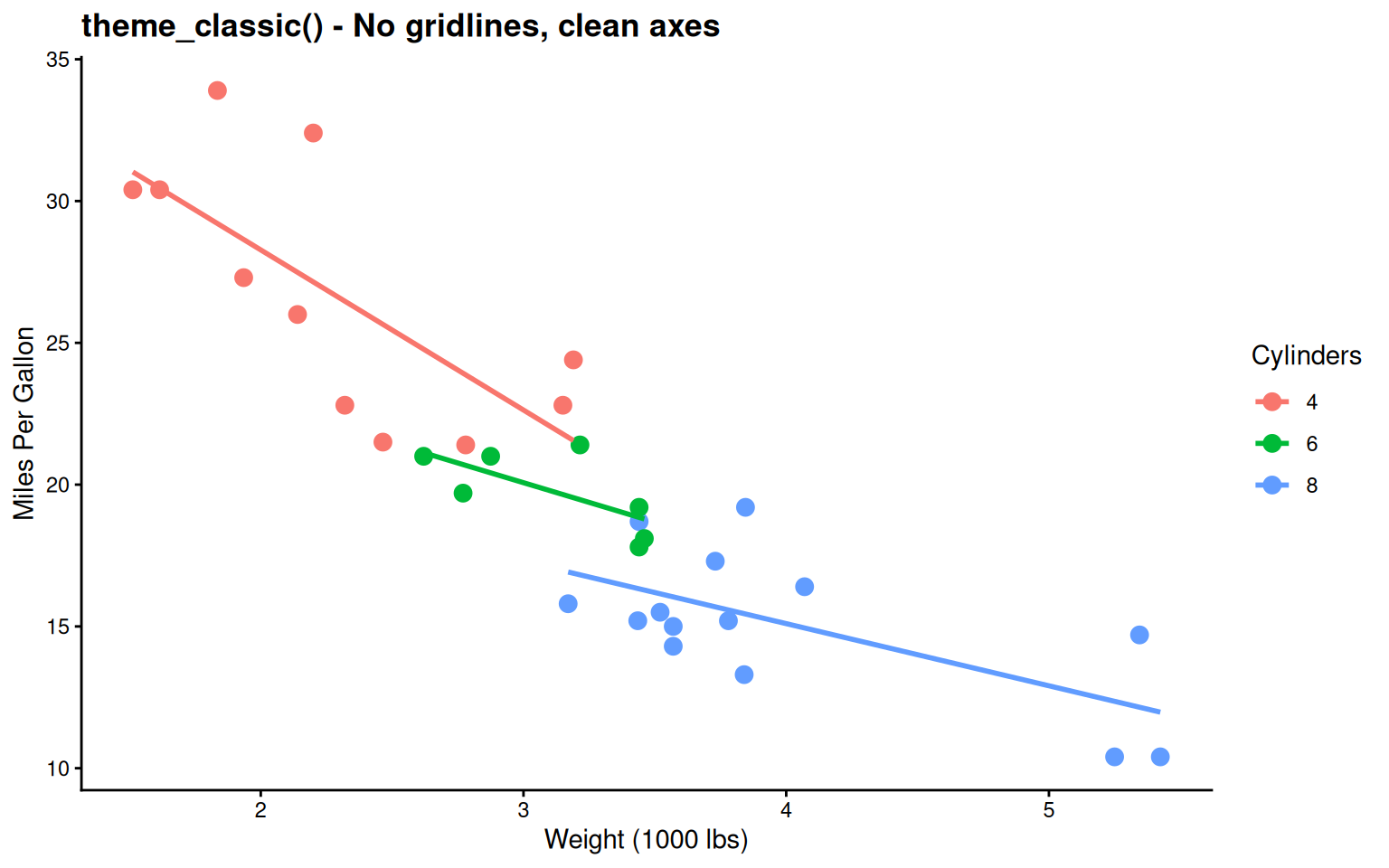
Very minimal - Traditional journal style
Built-in Theme: theme_minimal()
p <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE) +
labs(title = "theme_minimal() - Subtle gridlines, modern",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon",
color = "Cylinders") +
theme_minimal() +
theme(plot.title = element_text(face = "bold"))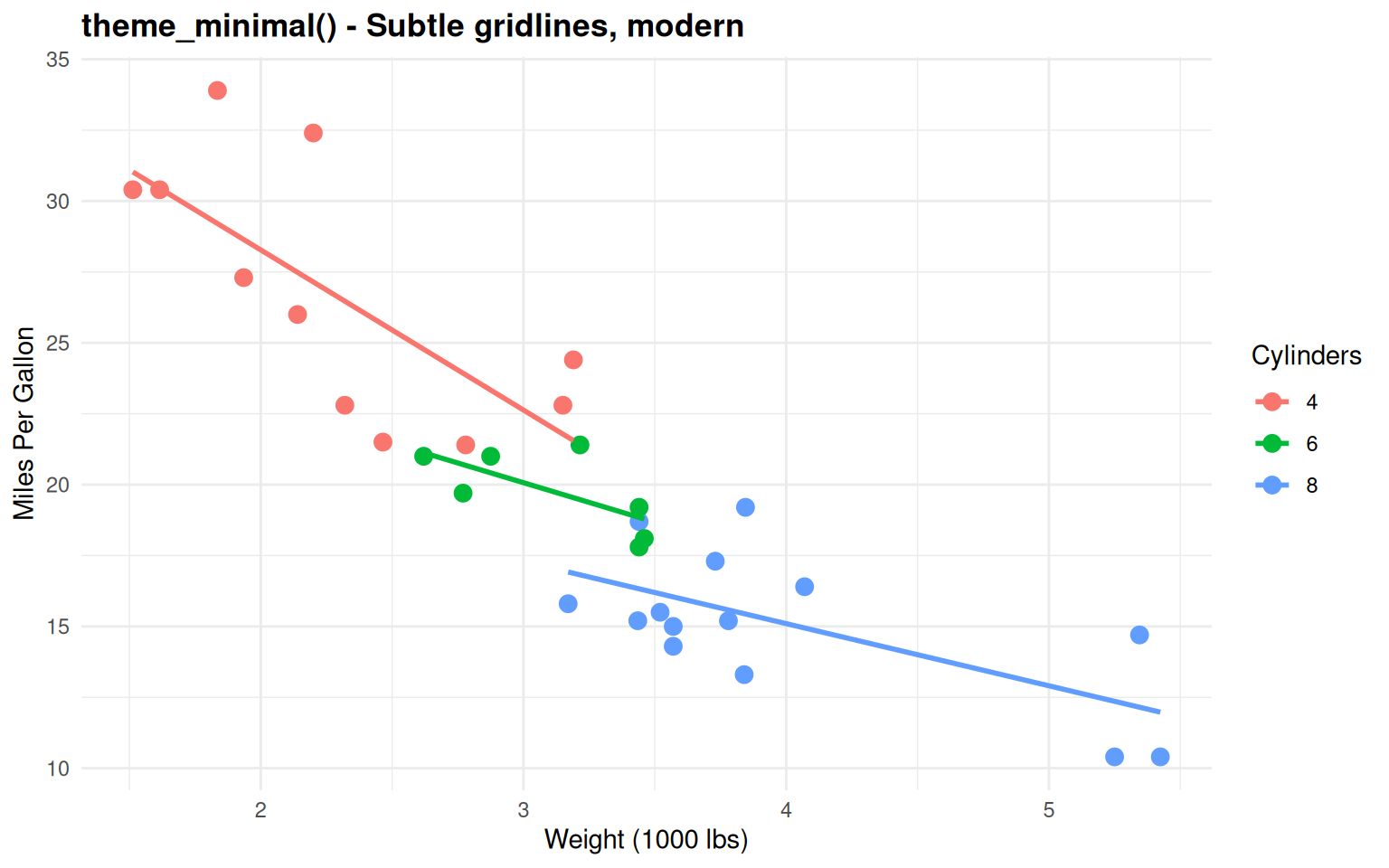
Good balance - Clean with subtle reference lines
Built-in Theme: theme_void()
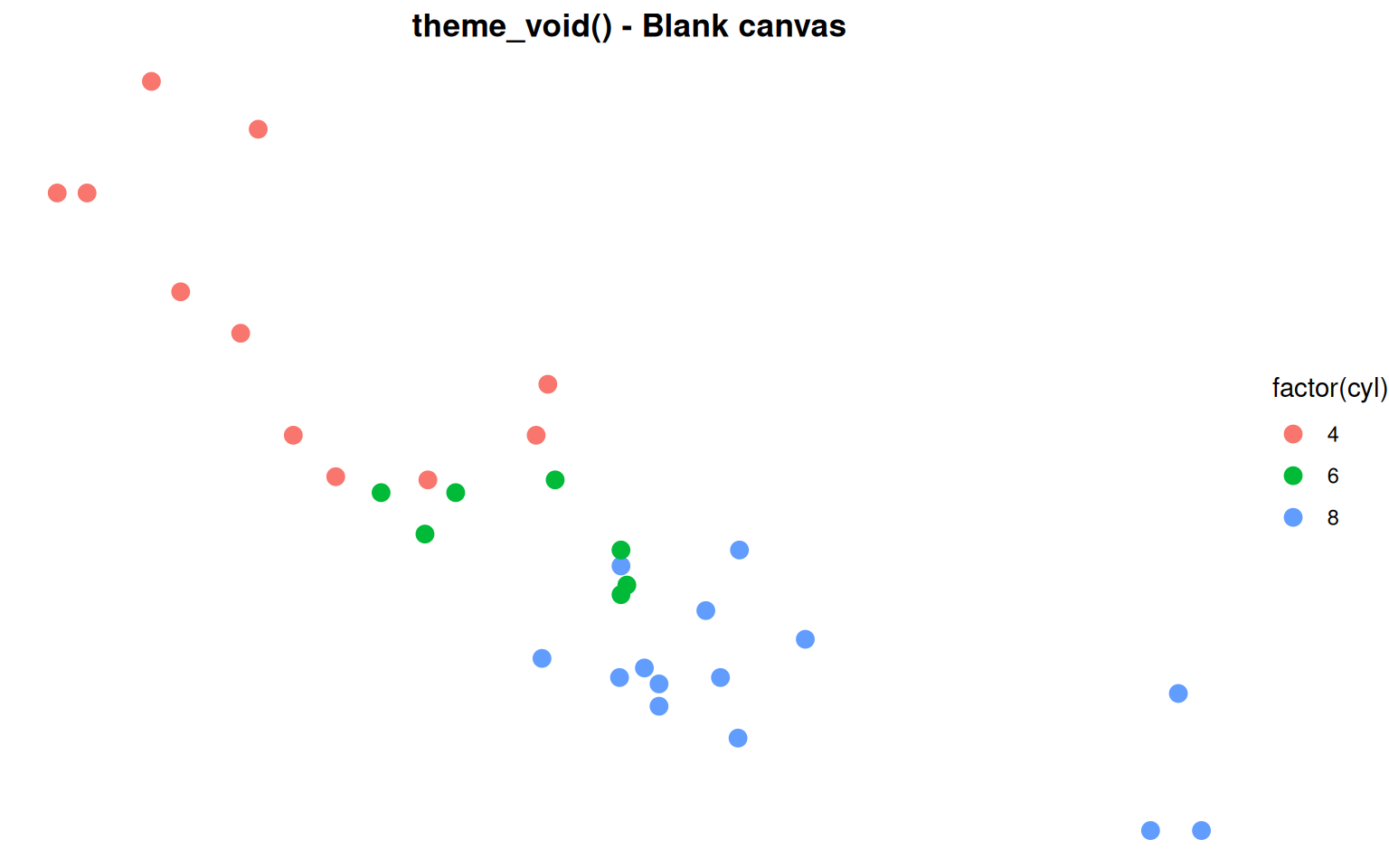
For custom designs - Maps, minimalist graphics
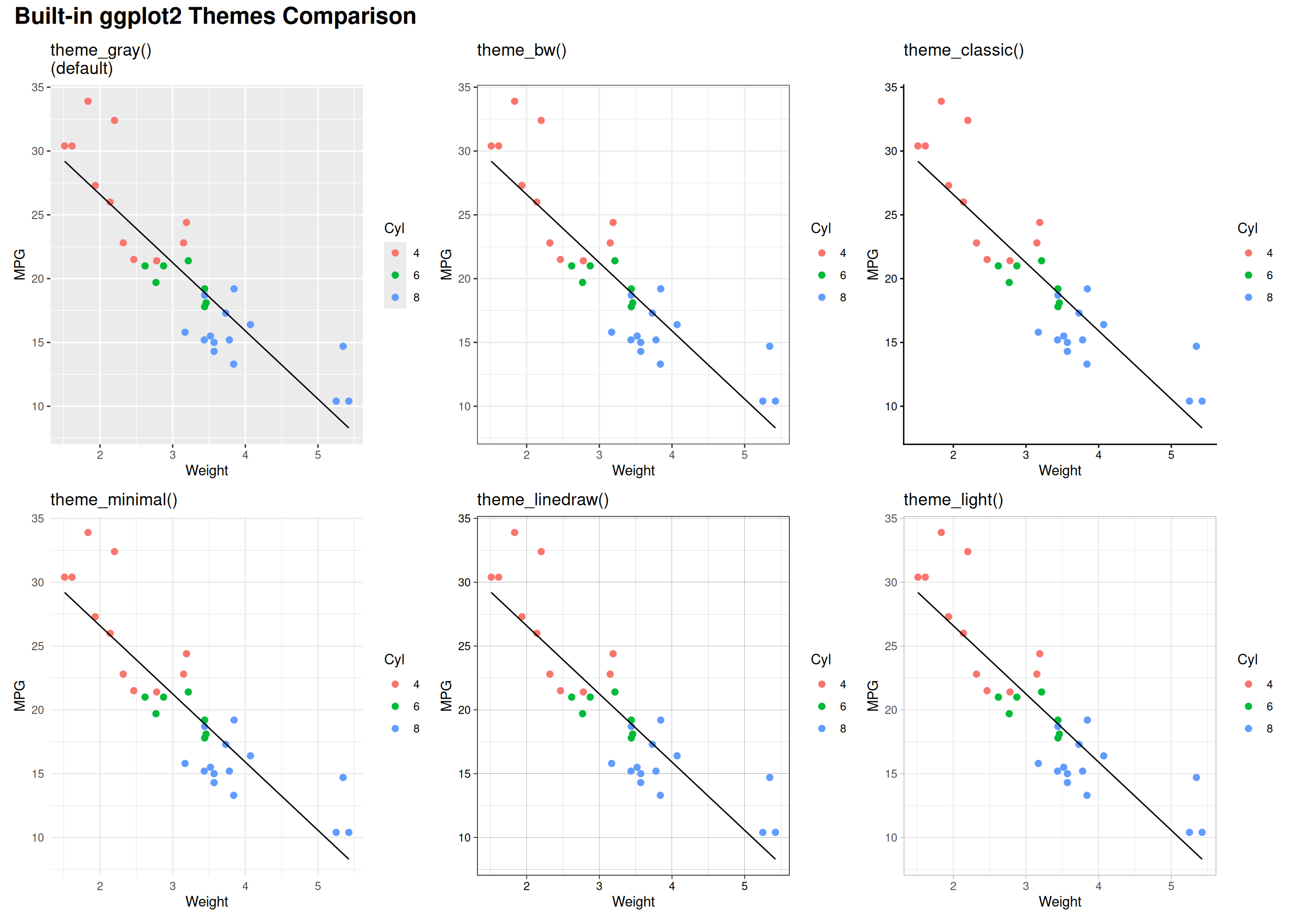
Side-by-Side Comparison
Publication Package: ggpubr
# ggpubr - publication-ready themes and statistical annotations
# install.packages("ggpubr")
ggplot(mtcars, aes(x = factor(cyl), y = mpg)) +
geom_boxplot(aes(fill = factor(cyl)), alpha = 0.7) +
geom_jitter(width = 0.2, alpha = 0.5) +
labs(title = "ggpubr - Publication ready with stats",
x = "Cylinders",
y = "Miles Per Gallon") +
theme_pubr() +
theme(legend.position = "none",
plot.title = element_text(face = "bold")) +
scale_fill_brewer(palette = "Set2")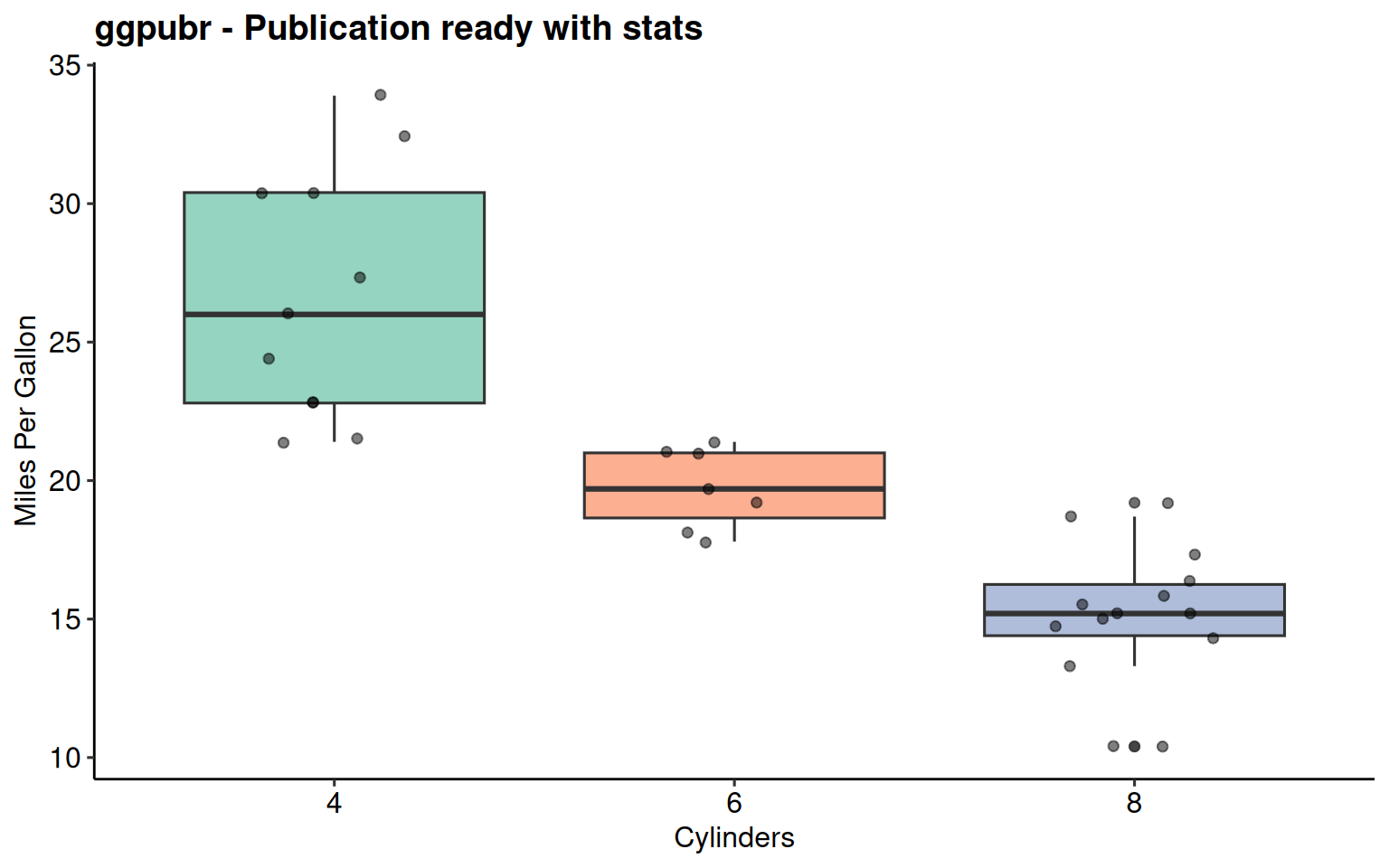
ggpubr Package
Publication-ready themes + statistical annotations
theme_pubr()- Clean publication themetheme_pubclean()- Even more minimalstat_regline_equation()- Automatic regression equationsstat_cor()- Correlation statisticsstat_compare_means()- p-values and significance brackets
ggpubr: Adding Regression Equations
library(ggpubr)
p <- ggplot(mtcars, aes(x = wt, y = mpg)) +
geom_point(size = 3, color = "steelblue") +
geom_smooth(method = "lm", se = TRUE,
color = "darkred", formula = y ~ x) +
stat_regline_equation(
aes(label = after_stat(eq.label)),
formula = y ~ x,
label.x.npc = 0.95, # 95% to the right (relative)
label.y.npc = 0.95, # 95% to the top (relative)
hjust = 1 # right-align text
) +
stat_cor(
aes(label = paste(after_stat(rr.label), after_stat(p.label), sep = "~~~~")),
label.x.npc = 0.95, label.y.npc = 0.88, hjust = 1
) +
labs(title = "Linear Regression with Equation",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon") +
theme_pubr()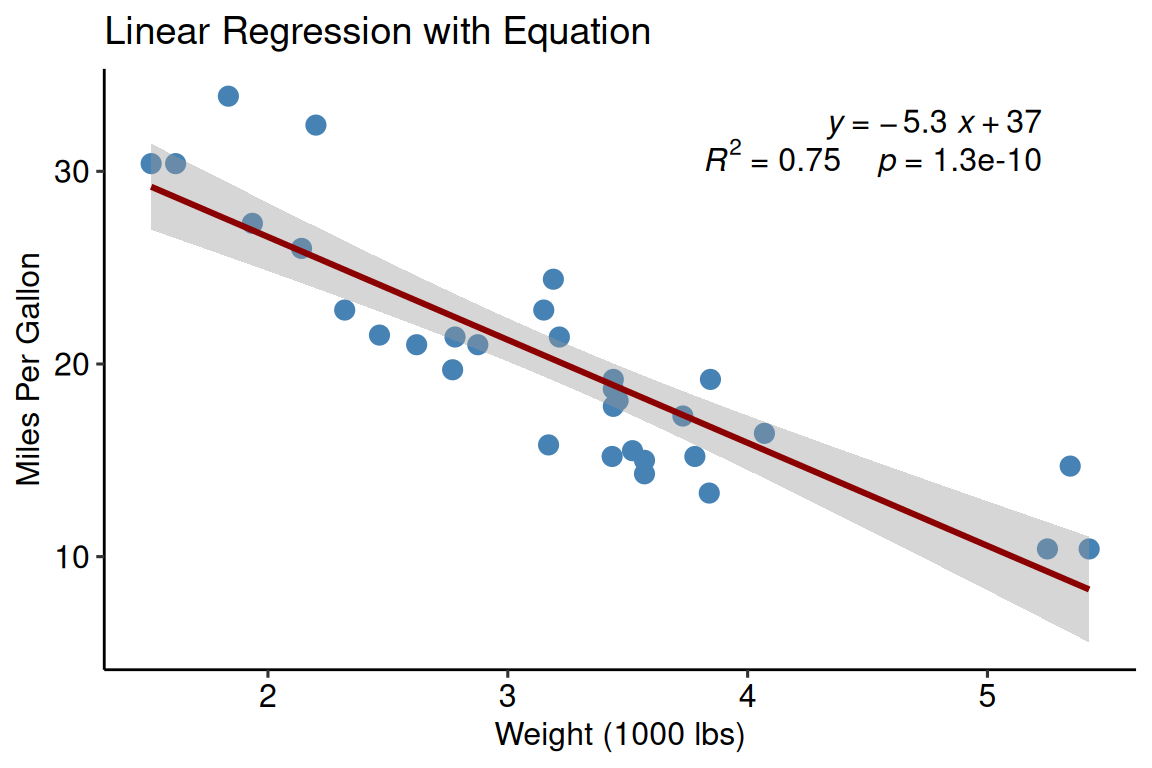
Key function: stat_regline_equation()
- Automatically calculates and displays equation
- Shows R² value
- Customizable position and formatting
- Works with facets
ggpubr: Pairwise Comparisons
# Automatically generate all pairwise comparisons
dose_levels <- levels(factor(ToothGrowth$dose))
my_comparisons <- combn(dose_levels, 2, simplify = FALSE)
p <- ggboxplot(ToothGrowth,
x = "dose", y = "len", color = "dose", palette = "jco") +
stat_compare_means(
comparisons = my_comparisons,
method = "t.test",
p.adjust.method = "BH" # Benjamini-Hochberg (FDR) correction
) +
stat_compare_means(
method = "anova",
label.y = 50
) +
labs(title = "Pairwise Comparisons with Multiple Testing Correction",
x = "Dose (mg/day)",
y = "Tooth Length")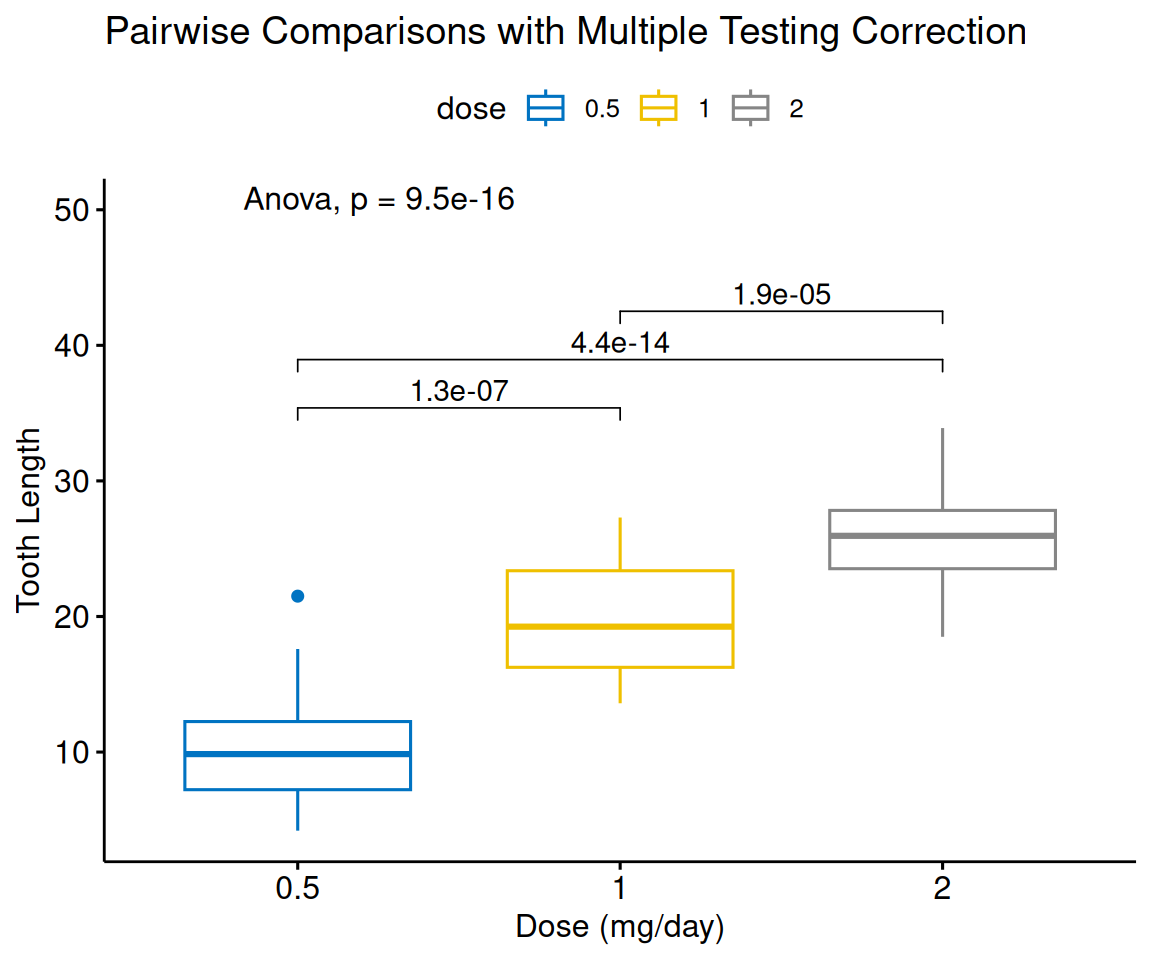
combn(levels, 2)generates all pairs automaticallymethod = "t.test"for pairwise tests (ormethod = "tukey_hsd"for Tukey’s HSD)p.adjust.method = "BH"for multiple testing correction (“holm”, “bonferroni”, “hochberg”, “BY”, “fdr”)method = "anova"for overall test- Automatic significance brackets
Modern Typography: hrbrthemes
ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE, formula = y ~ x) +
facet_wrap(~gear, labeller = label_both) +
labs(title = "hrbrthemes::theme_ipsum() - Modern typography",
subtitle = "Clean, professional, with excellent fonts and facets",
x = "Weight (1000 lbs)",
y = "Miles Per Gallon",
color = "Cylinders") +
theme_ipsum() +
scale_color_ipsum()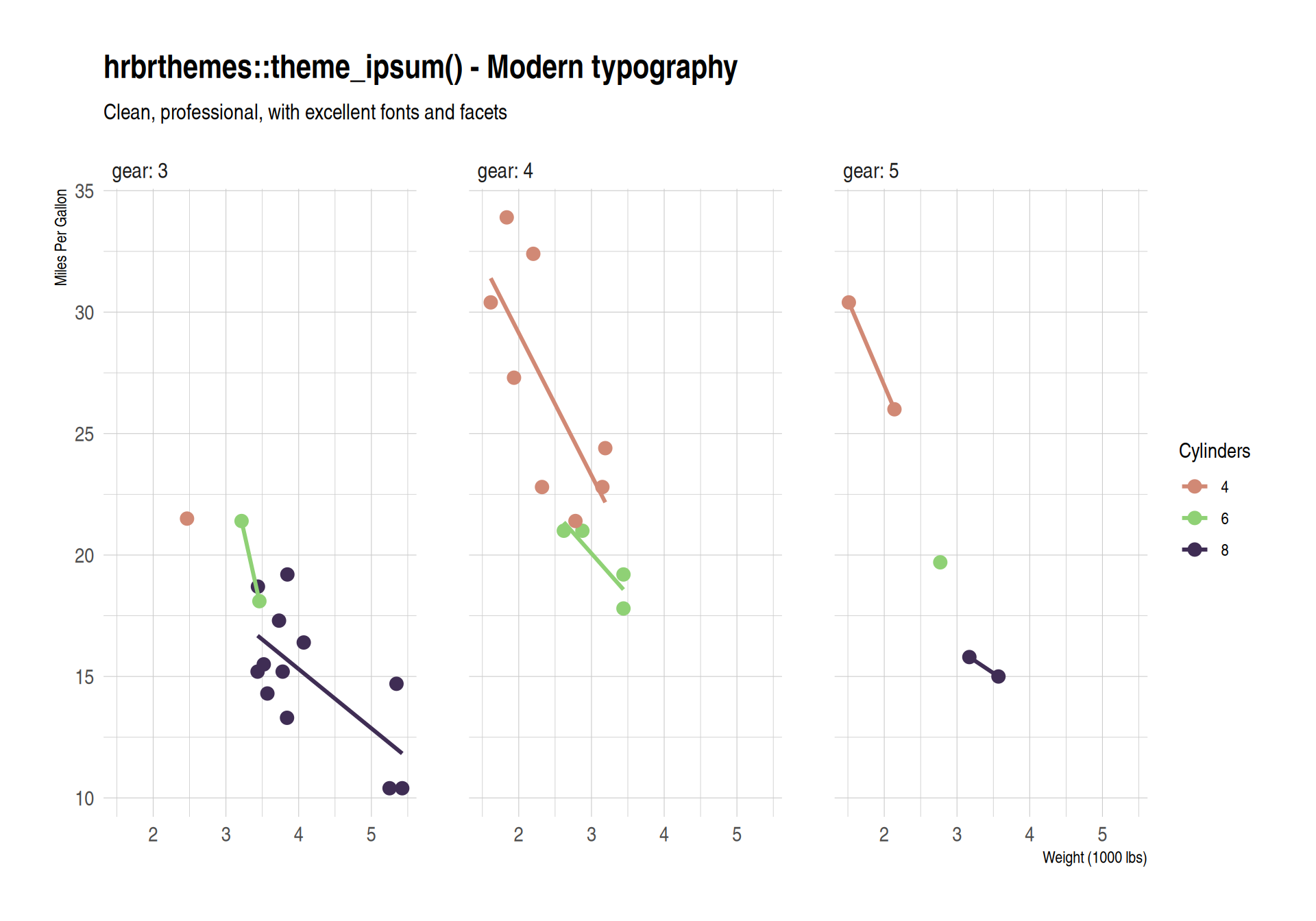
hrbrthemes Package
Modern professional typography
- Uses high-quality fonts (requires font installation)
theme_ipsum()- Modern, clean, professionaltheme_ipsum_rc()- Roboto Condensed font- Excellent for presentations and reports
- Works beautifully with facets
- May require:
extrafont::font_import()
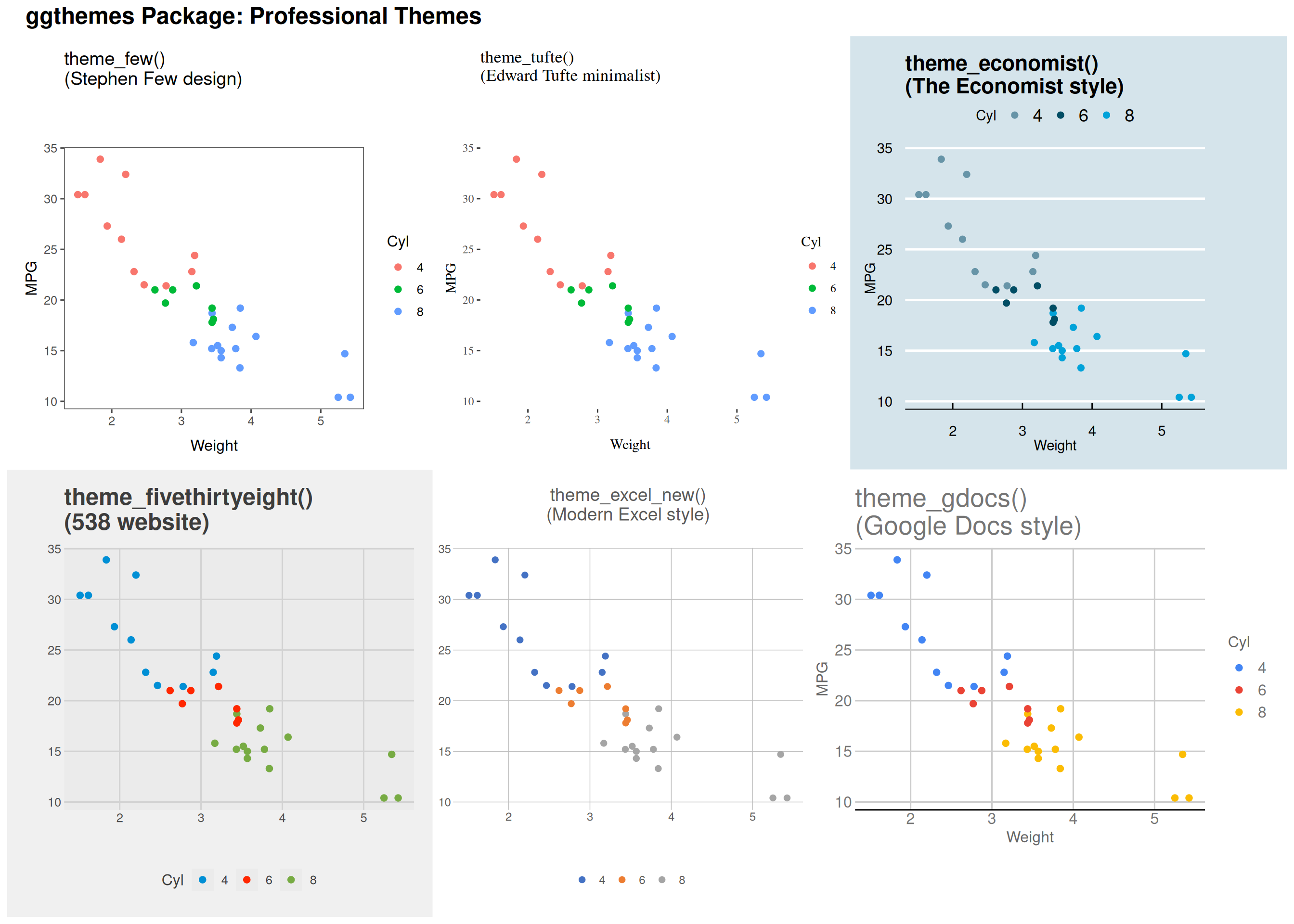
Specialized Themes: ggthemes
Setting Global Theme
Set once, apply to all plots:
# At top of script
theme_set(theme_bw(base_size = 12))
# Now all plots use theme_bw
# automatically
p1 <- ggplot(mtcars, aes(wt, mpg)) +
geom_point() +
labs(title = "Plot 1")
p2 <- ggplot(iris, aes(Sepal.Length, Sepal.Width)) +
geom_point(aes(color = Species)) +
labs(title = "Plot 2")
p3 <- ggplot(faithful, aes(eruptions)) +
geom_histogram(bins = 30, fill = "steelblue") +
labs(title = "Plot 3")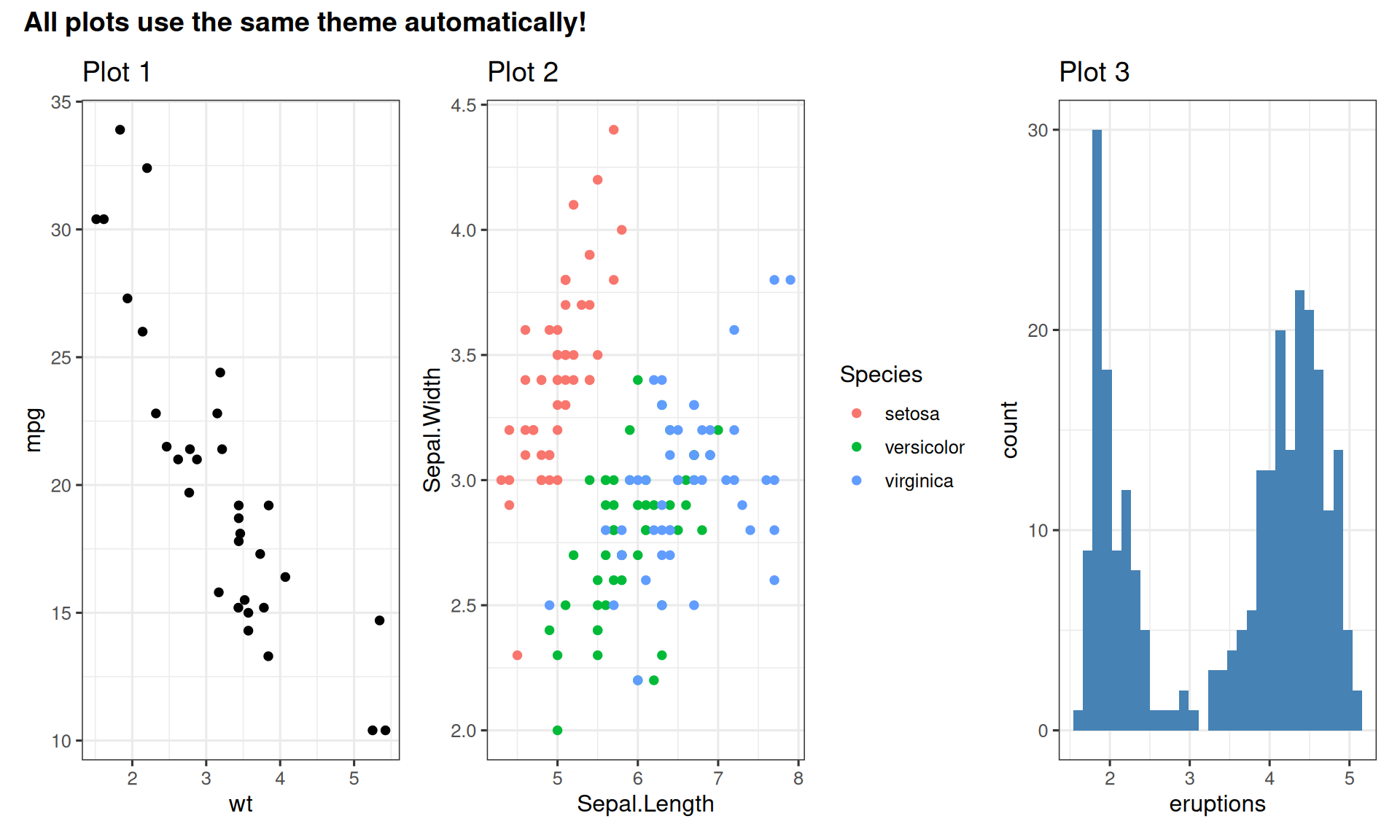
Font Considerations
Font Preferences by Journal
Many journals prefer specific fonts:
- Arial - Most common, widely accepted
- Helvetica - Classic choice
- Times New Roman - Traditional journals
- Calibri - Modern alternative
Check journal author guidelines!
Theme Elements You Can Customize
p <- ggplot(mtcars, aes(x = factor(cyl), y = mpg, fill = factor(cyl))) +
geom_boxplot(alpha = 0.7) +
facet_wrap(~gear, labeller = label_both) +
labs(title = "Customized Theme Elements",
x = "Cylinders",
y = "Miles Per Gallon",
fill = "Cylinders") +
theme_minimal() +
theme(
# Axis elements
axis.title = element_text(size = 12, face = "bold", color = "navy"),
axis.text = element_text(size = 10, color = "gray30"),
# Legend
legend.position = "bottom",
legend.title = element_text(face = "bold"),
legend.background = element_rect(fill = "gray95", color = "gray50"),
# Panel
panel.grid.major = element_line(color = "gray80", linewidth = 0.3),
panel.grid.minor = element_blank(),
panel.background = element_rect(fill = "white"),
# Facet strips
strip.background = element_rect(fill = "steelblue", color = "navy"),
strip.text = element_text(color = "white", face = "bold", size = 11),
# Plot
plot.title = element_text(size = 16, face = "bold", hjust = 0.5),
plot.background = element_rect(fill = "white", color = NA)
) +
scale_fill_brewer(palette = "Set2")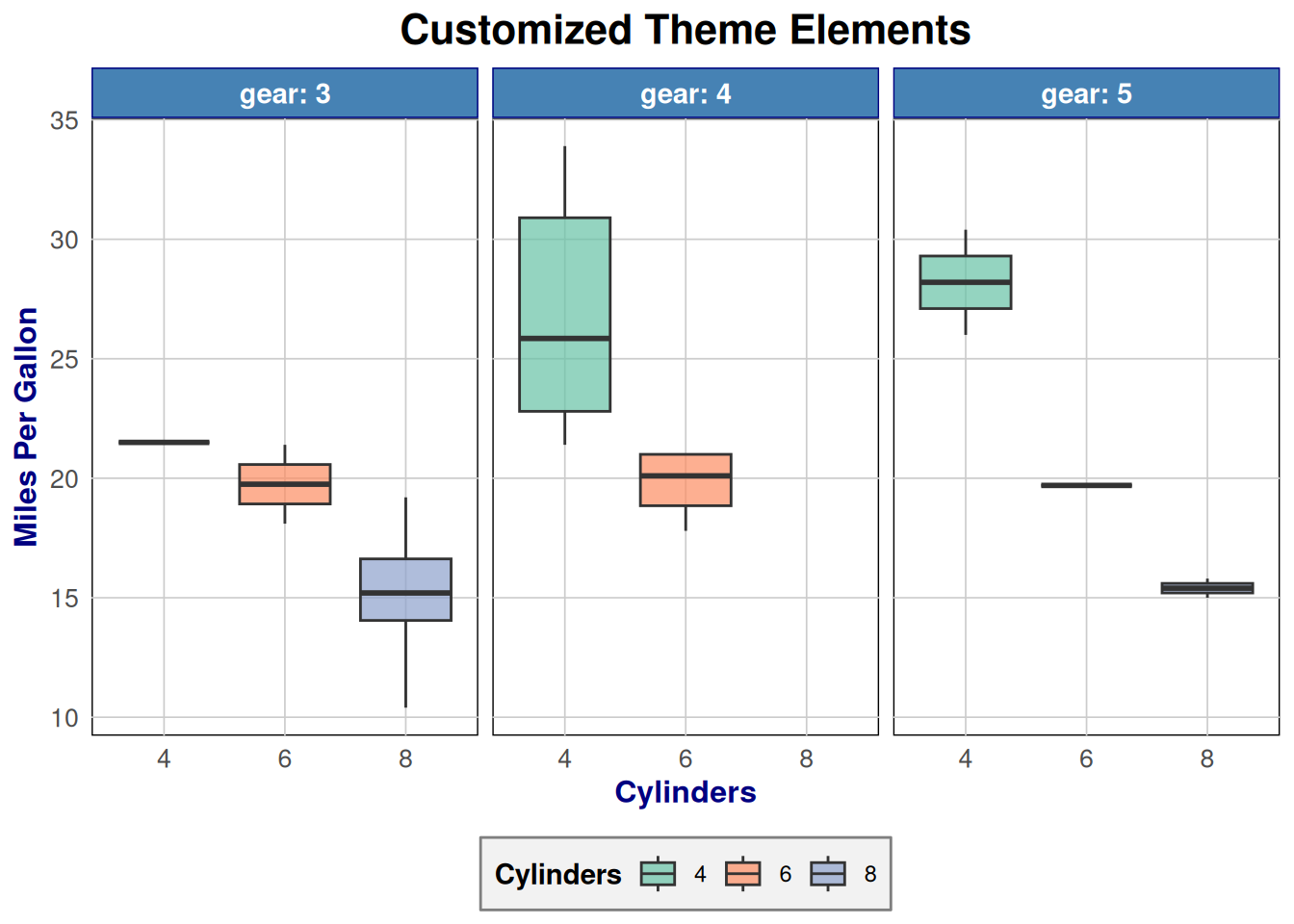
Customizable elements:
axis.title,axis.text- Axis labelslegend.position- “top”, “bottom”, “left”, “right”, “none”panel.grid- Gridlinesplot.background,panel.background- Backgroundsstrip.background,strip.text- Facet labels
Recommendations
Best Practices
- Never use default
theme_gray()for publications- Gray background wastes ink and looks unprofessional
- Set global theme at start of script for consistency
theme_set(theme_classic())applies to all subsequent plots
- Match journal style - check published figures
- Look at recent issues of your target journal
- Note font choices, gridline presence, color schemes
- Keep it simple - less chart junk = better
- Remove unnecessary gridlines
- Minimize non-data ink
- Use publication packages for multi-panel figures
patchworkfor intuitive combining syntaxggpubr::ggarrange()for automatic labeling
Saving Plots in R
The Graphics Device System
Every plot needs a “device”
Device = where R sends the graphics output (device = “the printer”)
- Screen (RStudio viewer)
- PDF file
- SVG file
- PNG file
- JPEG file
- etc.
Old Way: Manual Device Management
dev.off() Works with ALL R Plots
The dev.off() approach works for:
- Base R plots (
plot(),hist(),barplot(), etc.) - ggplot2 plots
- Any R graphics output
It’s universal - not limited to any specific plotting system!
Problems with manual device management:
- Easy to forget
dev.off() - Verbose
- Not intuitive
- Must manage device lifecycle manually
Modern Way: ggsave()
For ggplot2 objects (recommended!)
No dev.off() needed! ✨
Full Control
Batch Saving: Result
# A tibble: 3 × 3
Species data plot
<fct> <list> <list>
1 setosa <tibble [50 × 4]> <gg>
2 versicolor <tibble [50 × 4]> <gg>
3 virginica <tibble [50 × 4]> <gg>Each row contains:
- Species name
- Nested data for that species
- A ggplot object with regression
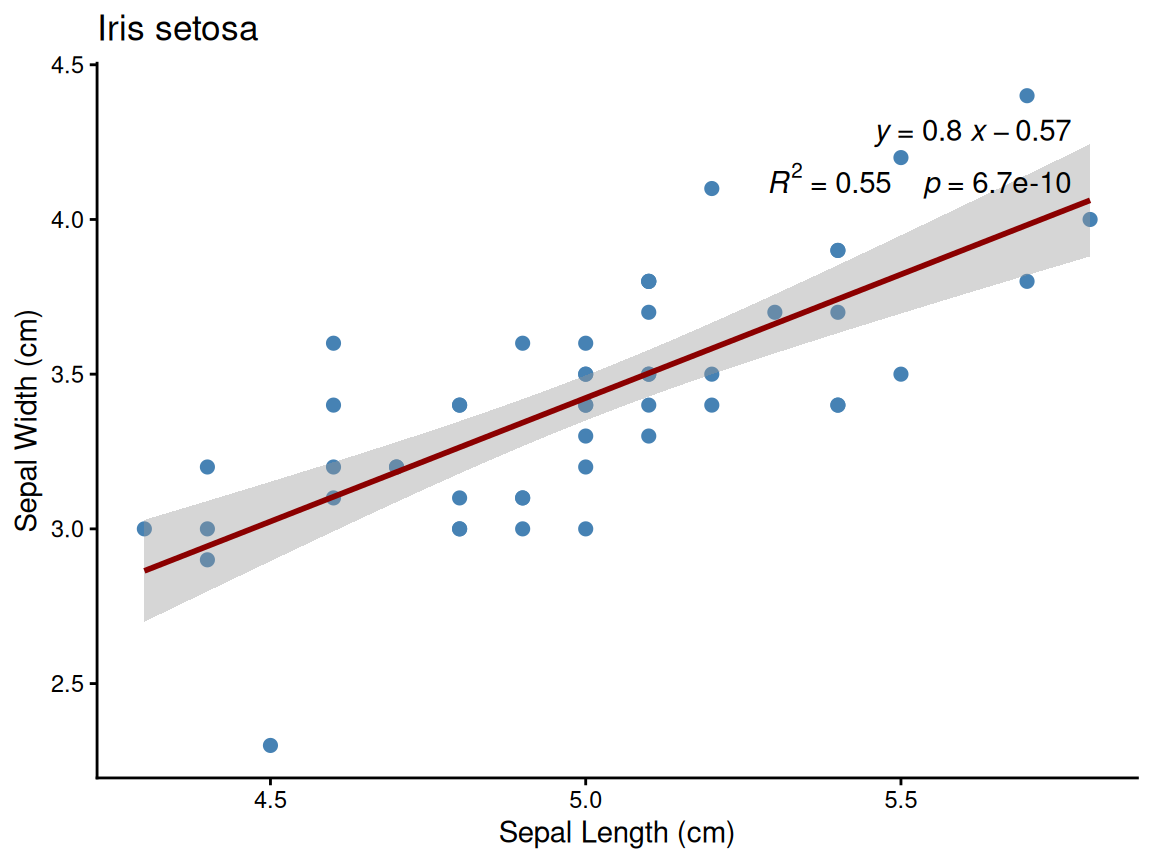
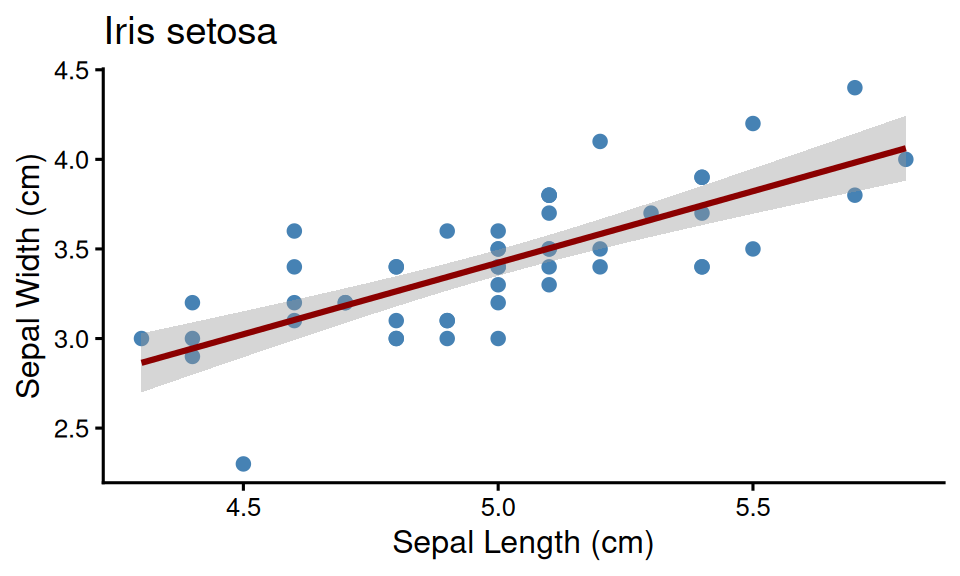
Example: Setosa species plot
File Organization
Using magick for Conversion
library(magick)
# Convert PDF to 300 DPI PNG
img <- image_read_pdf("plot.pdf", density = 300)
image_write(img, "plot.png", format = "png", quality = 100)
# With better antialiasing (remove alpha channel)
img <- image_read_pdf("plot.pdf", density = 300)
img <- image_background(img, "white") # Remove alpha
image_write(img, "plot.png", format = "png", quality = 100)
# Batch convert all PDFs in directory
pdf_files <- list.files(pattern = "\\.pdf$")
for (file in pdf_files) {
img <- image_read_pdf(file, density = 300)
img <- image_background(img, "white")
out_file <- sub("\\.pdf$", ".png", file)
image_write(img, out_file, format = "png", quality = 100)
}Batch Saving Multiple Plots
Make a plotting function ::: {.cell}
make_species_plot <- function(data, species) {
ggplot(data, aes(x = Sepal.Length, y = Sepal.Width)) +
geom_point(size = 2, color = "steelblue") +
geom_smooth(method = "lm", se = TRUE, color = "darkred", formula = y ~ x) +
labs(title = glue("Iris {species}"), x = "Sepal Length (cm)", y = "Sepal Width (cm)") +
theme_classic(base_size = 12)
}
Nest data per Species ::: {.cell}
Write out to separate file per Species
::::
# A tibble: 3 × 3
Species data plot
<fct> <list> <list>
1 setosa <tibble [50 × 4]> <ggplt2::>
2 versicolor <tibble [50 × 4]> <ggplt2::>
3 virginica <tibble [50 × 4]> <ggplt2::># A tibble: 6 × 4
Sepal.Length Sepal.Width Petal.Length Petal.Width
<dbl> <dbl> <dbl> <dbl>
1 5.1 3.5 1.4 0.2
2 4.9 3 1.4 0.2
3 4.7 3.2 1.3 0.2
4 4.6 3.1 1.5 0.2
5 5 3.6 1.4 0.2
6 5.4 3.9 1.7 0.4
::::
Recommendations
Best Practices
- Always use ggsave() for ggplot2 (not manual devices)
- Always specify width, height, units and DPI (for raster output)
- Increase DPI from the default for all raster formats set ≥300
- Use cairo_pdf for PDF output
- Save both PDF and PNG versions
- Organize figures in dedicated directory
Post-Export Editing
Why Edit After Export?
Legitimate uses:
- Combine multiple plots into multi-panel figures (A, B, C labels)
- Fine-tune alignment and spacing
- Add annotations, arrows, or highlights
- Adjust layout without re-running analysis
Vector Editing Tools
Inkscape
- Free & open source
- Native SVG format
- Excellent PDF import
- Cross-platform
- Full-featured vector editor
- Converts between vector formats
- Slow…
Other options:
- Adobe Illustrator: Industry standard (commercial)
- PowerPoint: Can edit SVG as vector format
⚠️ Avoid Raster Editors!
DO NOT use Photoshop, GIMP, or other raster editors for plots!
- These convert your plots to pixels (rasterization)
- You lose scalability and editability
- Text becomes uneditable
- Quality degrades when resized
Keep it vector! Use Inkscape, Illustrator, or PowerPoint with SVG input and PDF output.
Inkscape Basics
Opening PDFs/SVGs:
- File → Open → Select PDF or SVG
- Each plot element is now editable
Useful tools:
- Selection tool (F1): Move and resize
- Text tool (F8): Edit or add text
- Align and Distribute (Ctrl+Shift+A)
- Guides (drag from rulers): Align elements precisely
Tips:
- Group related elements (Ctrl+G)
- Lock layers to prevent accidental edits
- Use layers for complex figures
Handling Missing Fonts:
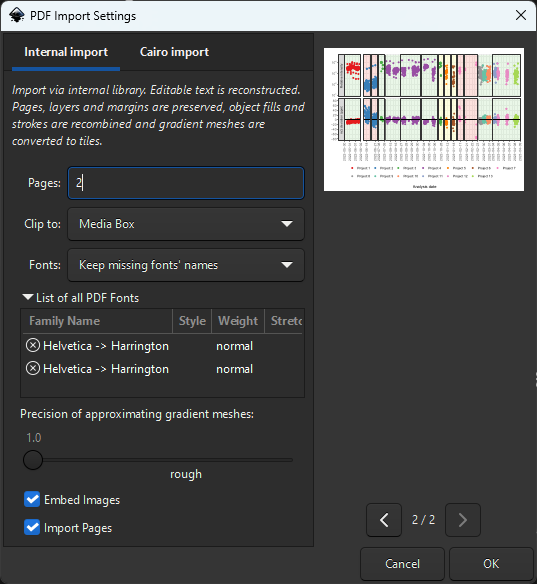
Keep the font names! Don’t substitute - preserves original font info and prevents text reflow issues
Cropping Canvas to Remove Whitespace
The Page Tool approach:
- Select the objects you want to keep (or Select All (Ctrl+A))
- Use Edit → Resize Page to Selection (or Ctrl+Shift+R)
- This resizes the canvas to fit your selection
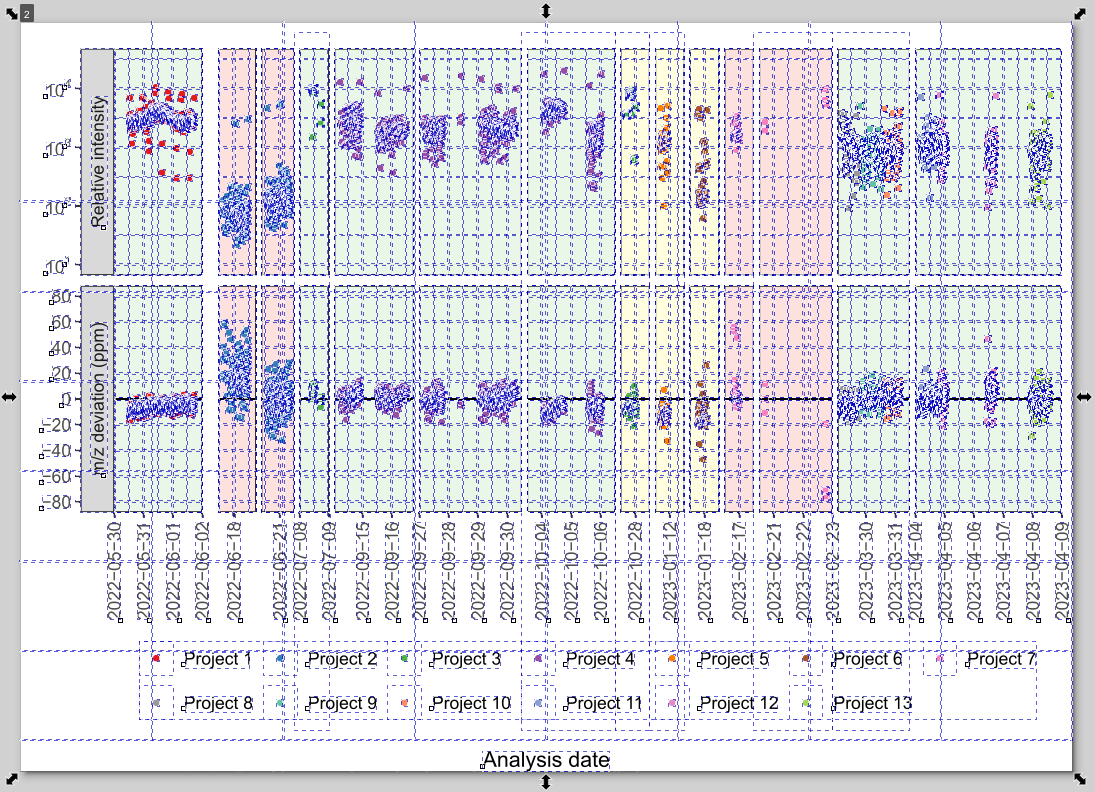
Hidden Objects from R Plots!
R exports contain many invisible/empty objects that prevent proper cropping!
The frustrating whack-a-mole:
- Press Ctrl+A (Select All) to reveal hidden objects
- You’ll see many white/empty rectangles
- Delete these empty objects first before resizing page
- Otherwise canvas includes invisible whitespace
Why clipping doesn’t work:
- Object → Clip → Set can destroy plot elements
- Don’t use clipping - delete empty objects instead
Example: Before and After Editing
Original R output
After editing in Inkscape
Changes made:
- Cropped whitespace
- Added annotations
- Made legend more compact
Combine figures directly in R instead
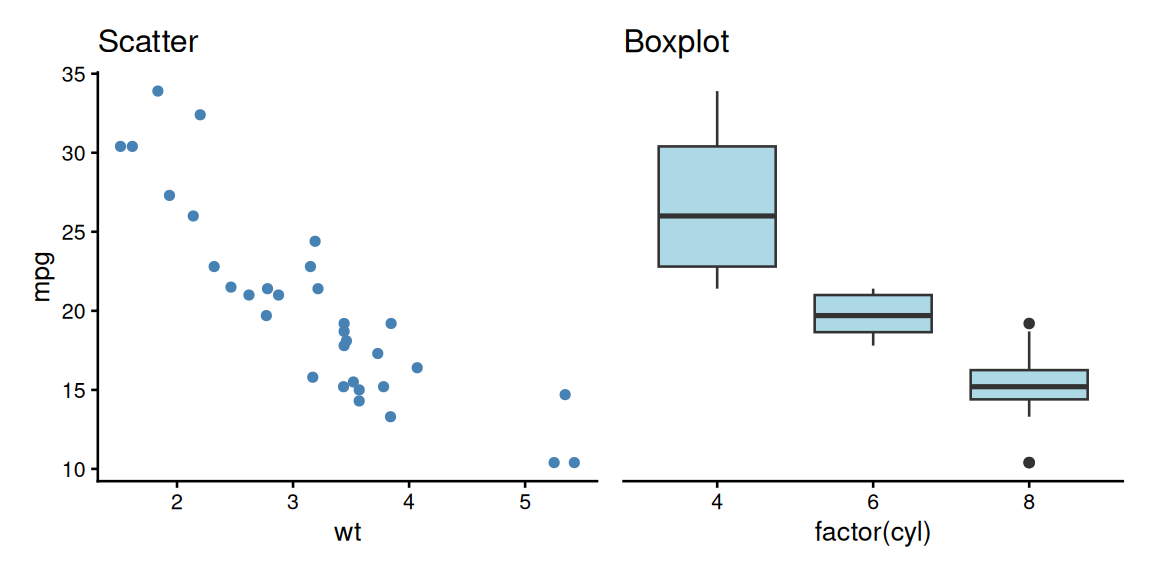
patchwork: Side by Side
A better alternative to manual composition!
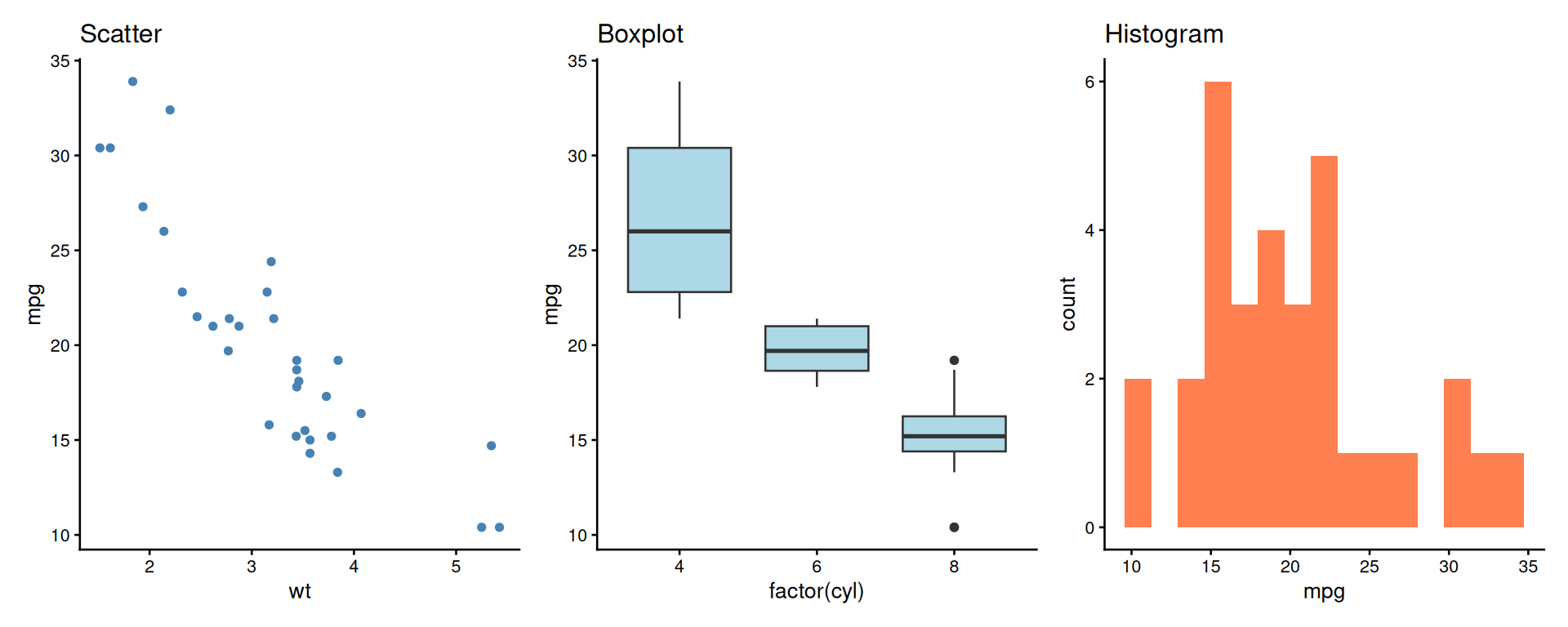
patchwork: Stacked Layout
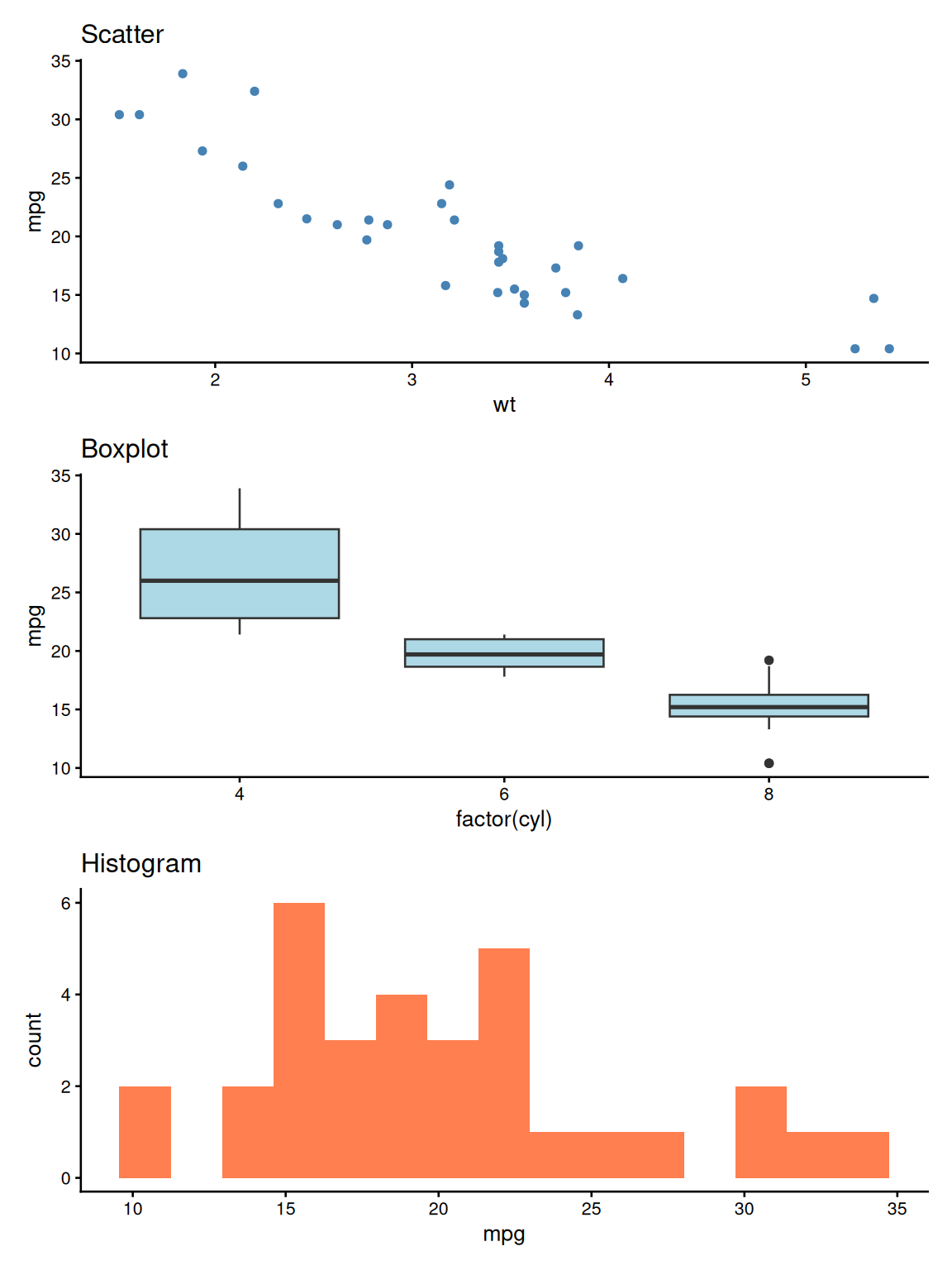
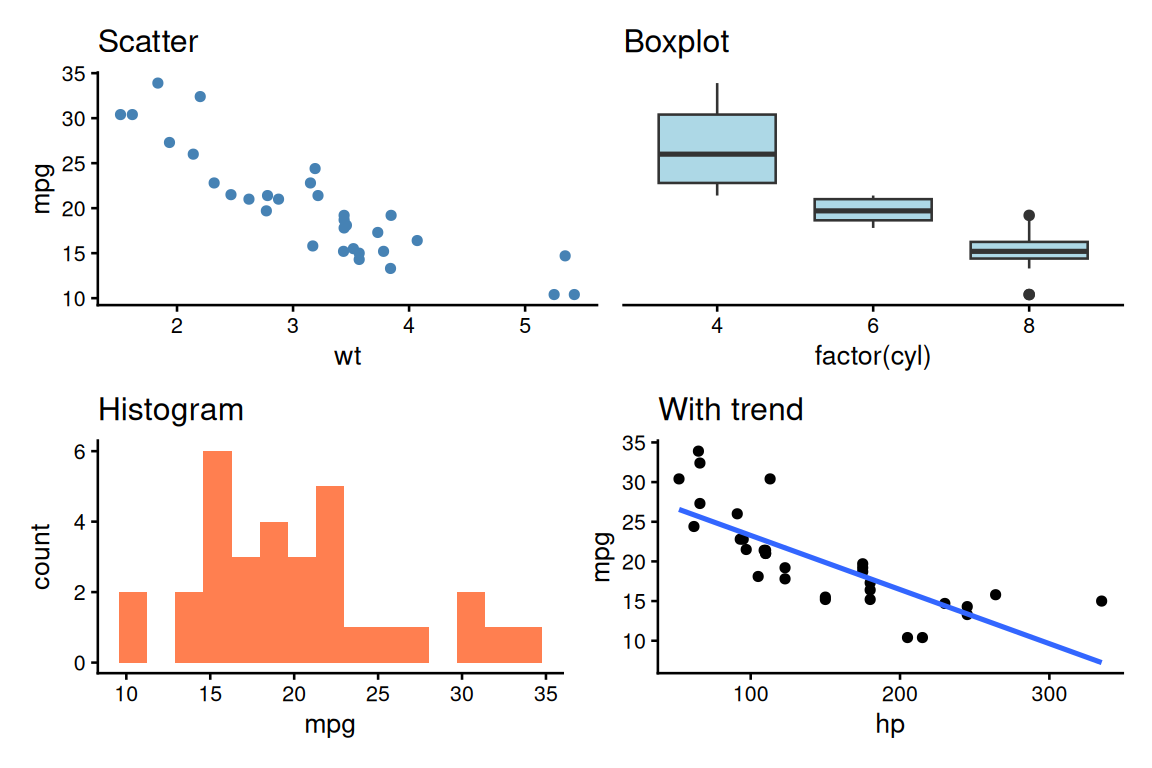
patchwork: Grid Layout
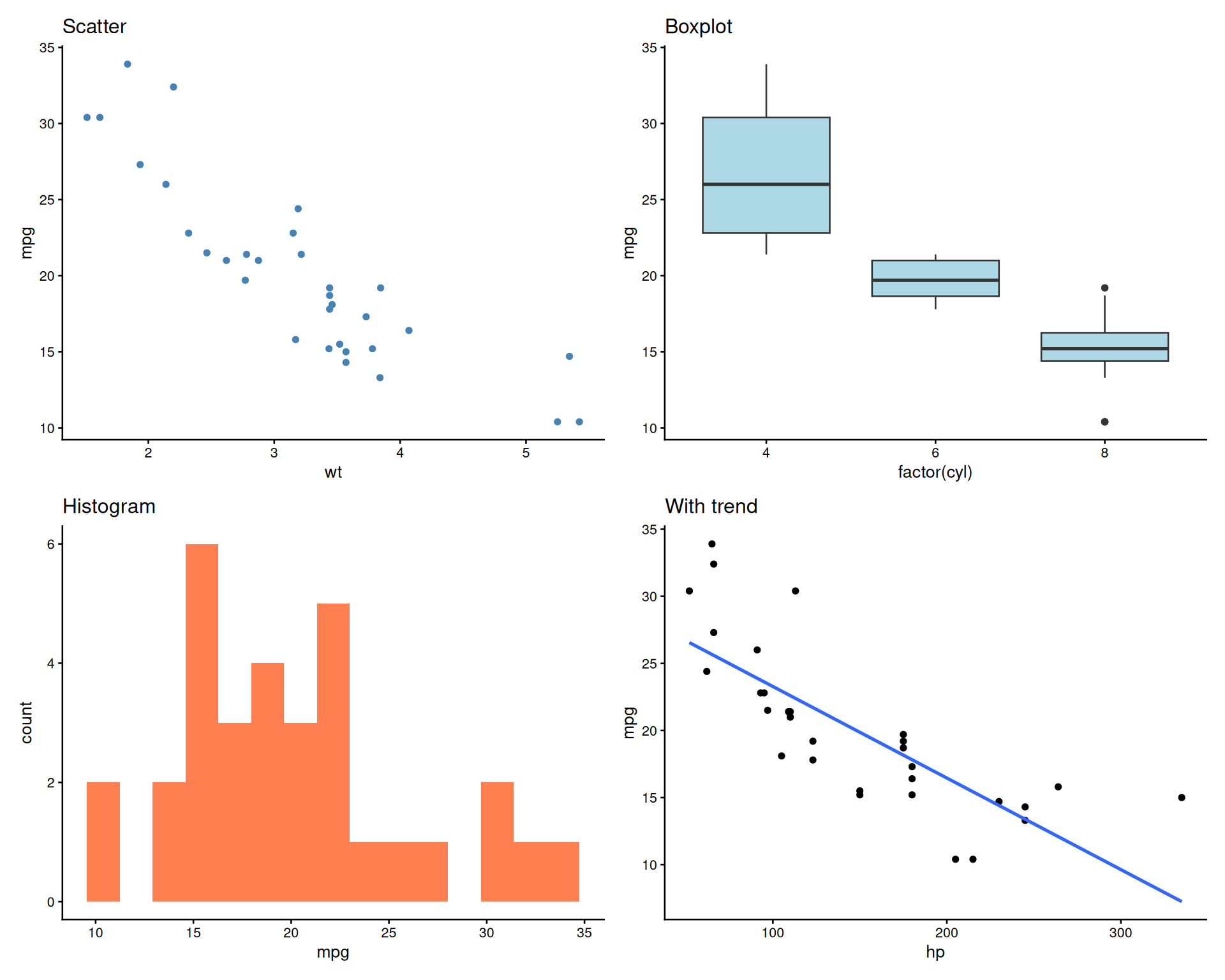
Adding Panel Labels (A, B, C…)
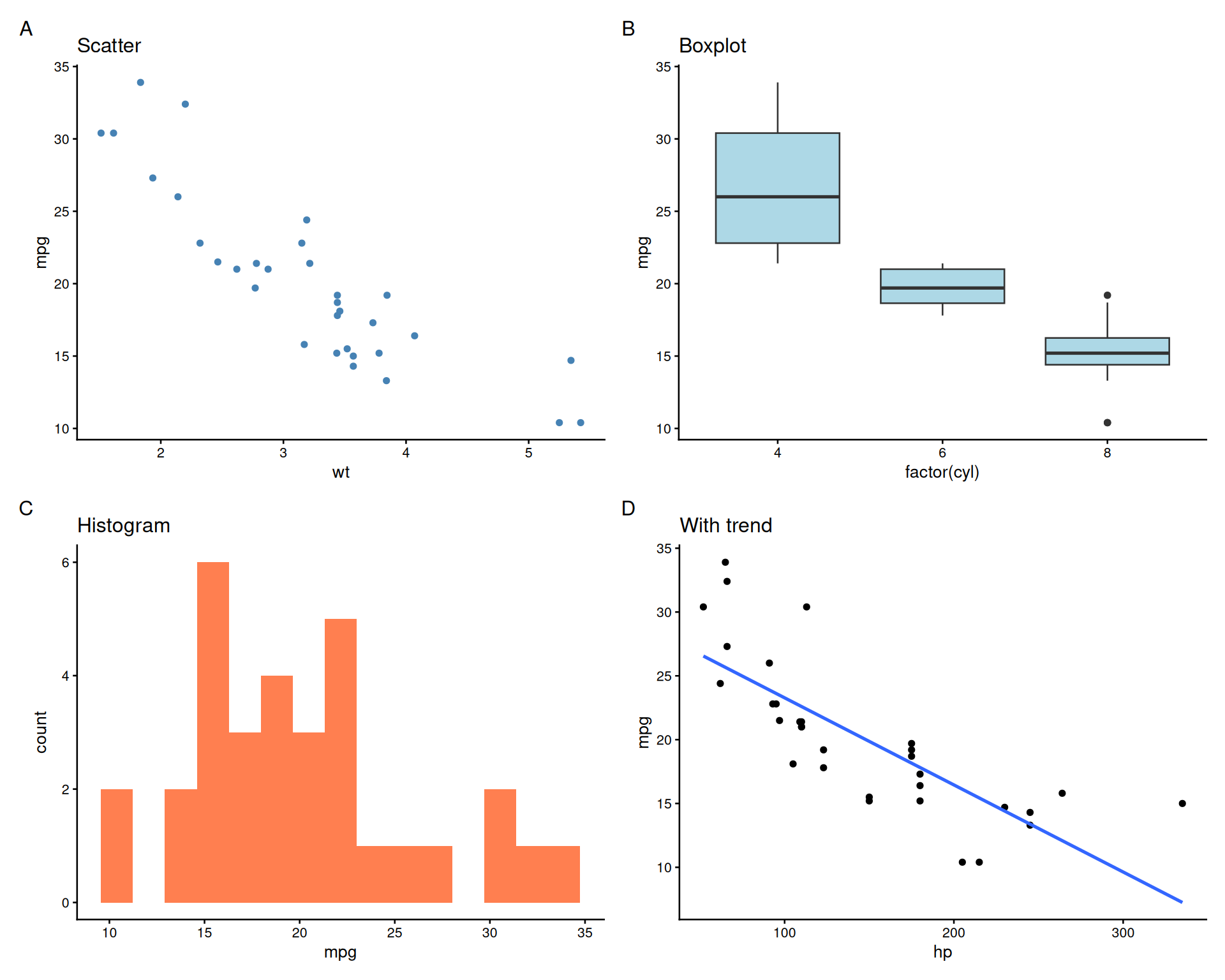
Customizing Panel Labels
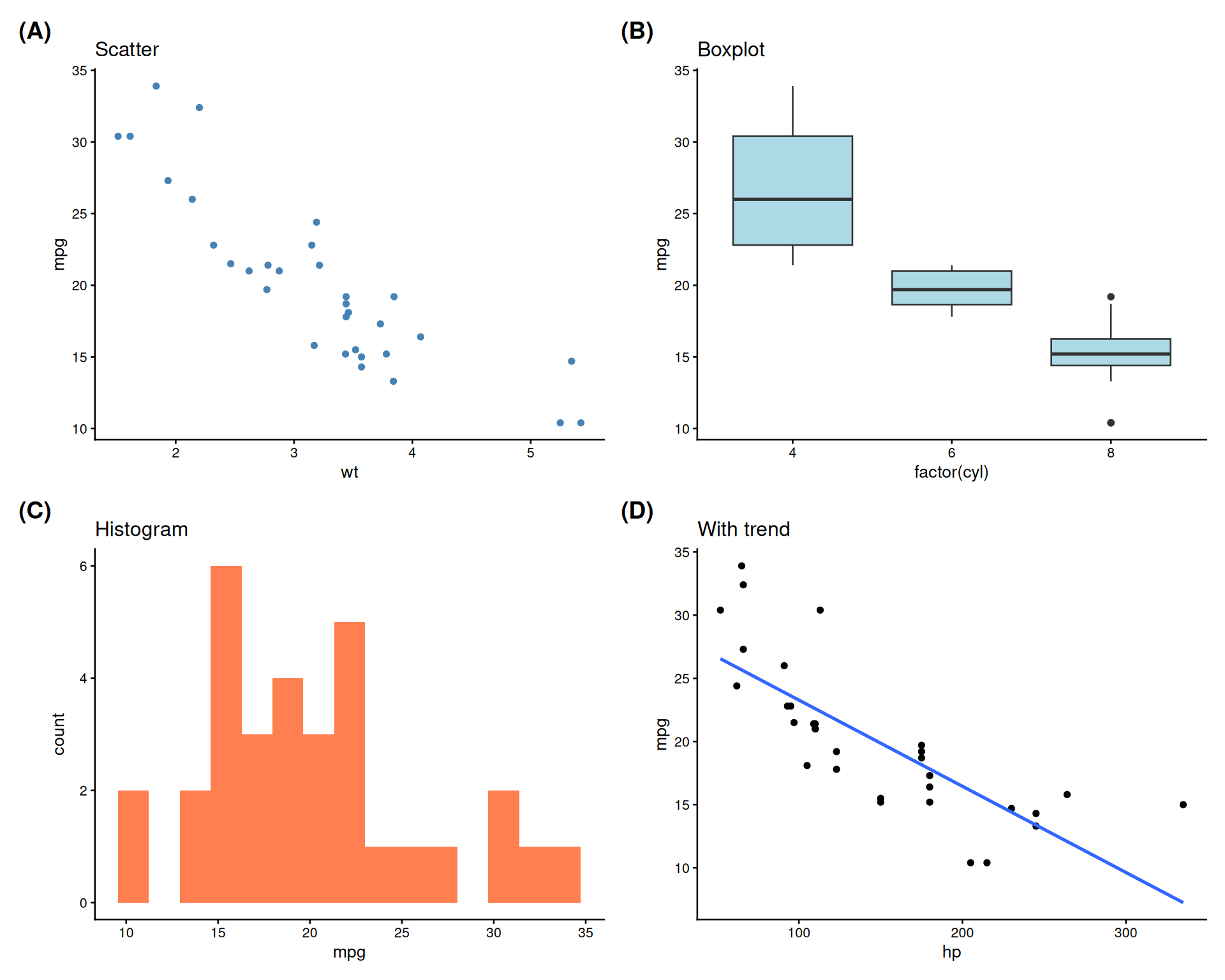
Unequal Panel Sizes
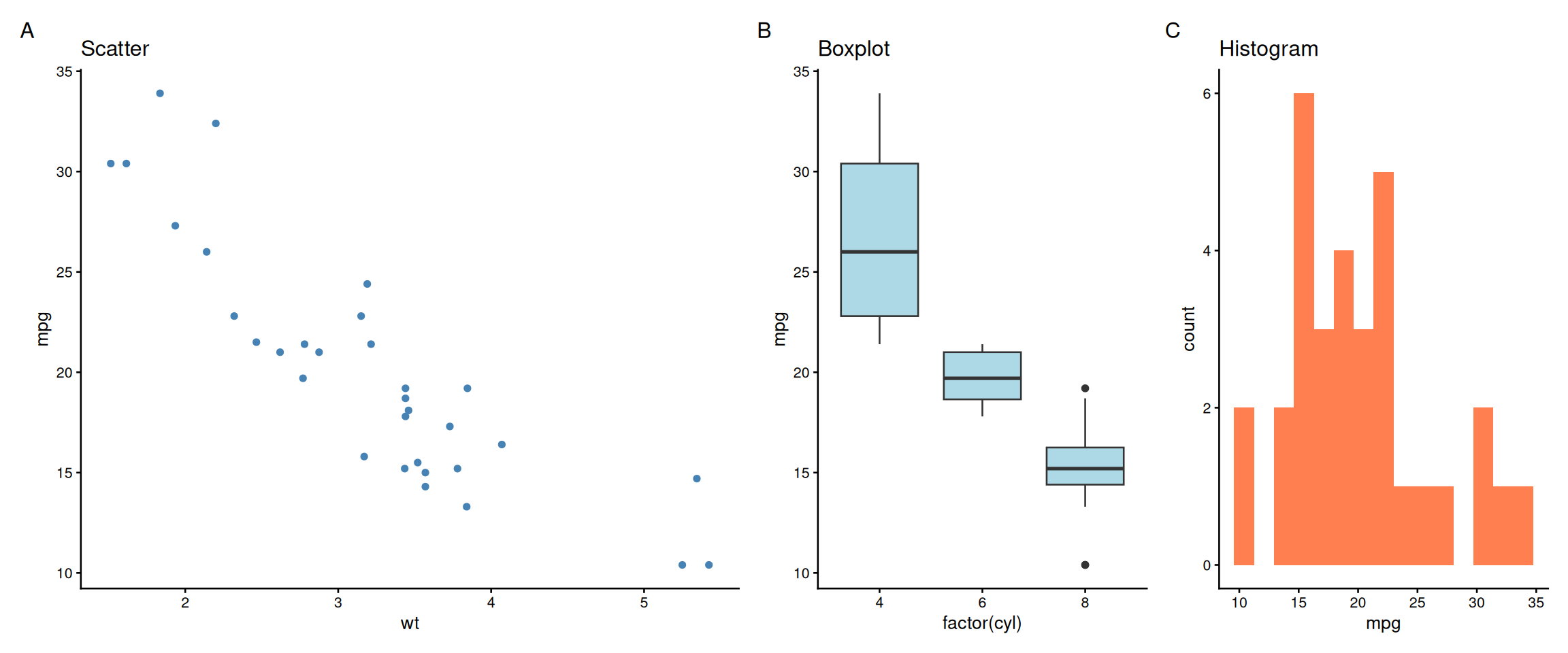
Nested Layouts
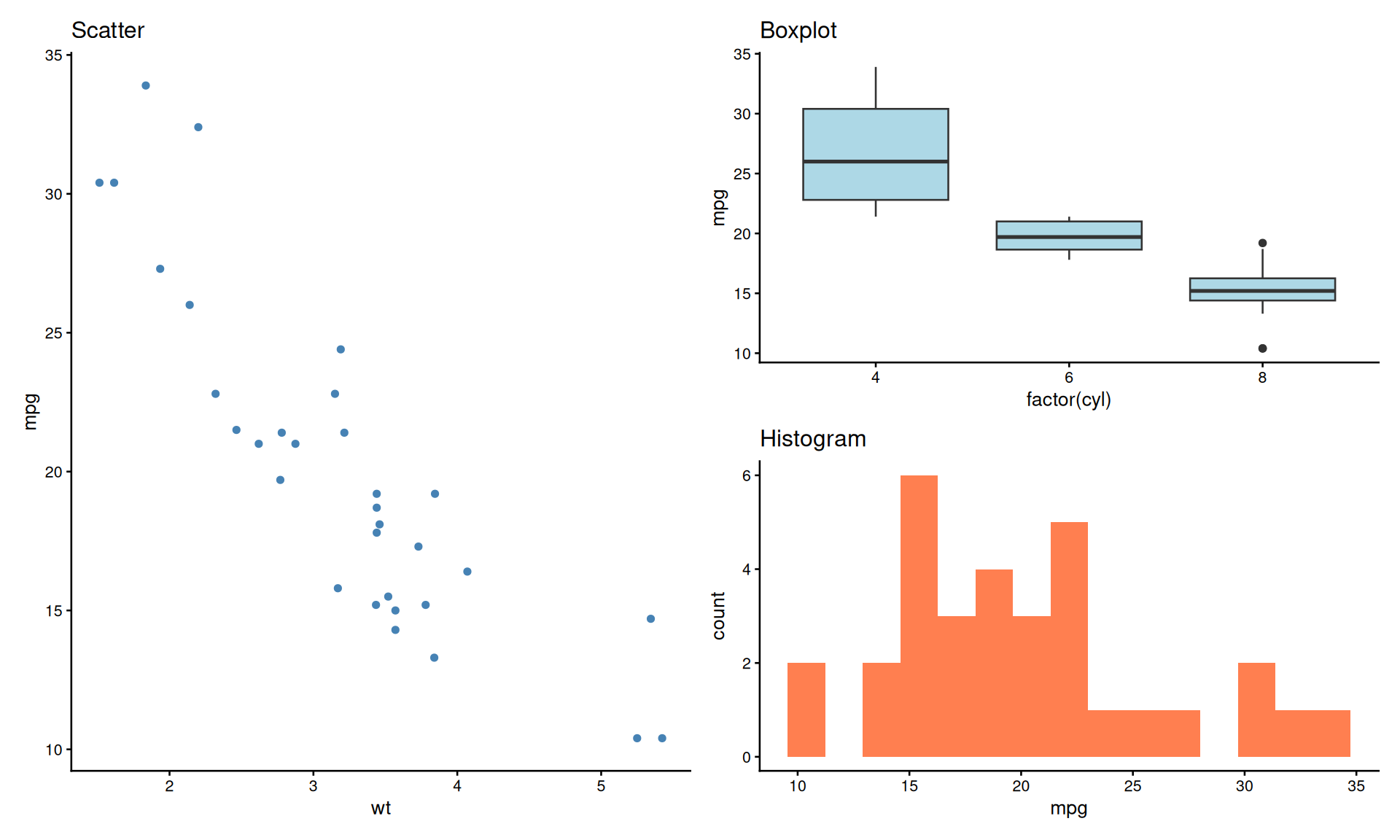
Shared Legends with plot_layout()
# Create plots with same color mapping
pa <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point() + theme_classic(base_size = 10)
pb <- ggplot(mtcars, aes(hp, mpg, color = factor(cyl))) +
geom_point() + theme_classic(base_size = 10)
pa + pb +
plot_layout(guides = 'collect') +
plot_annotation(tag_levels = 'A')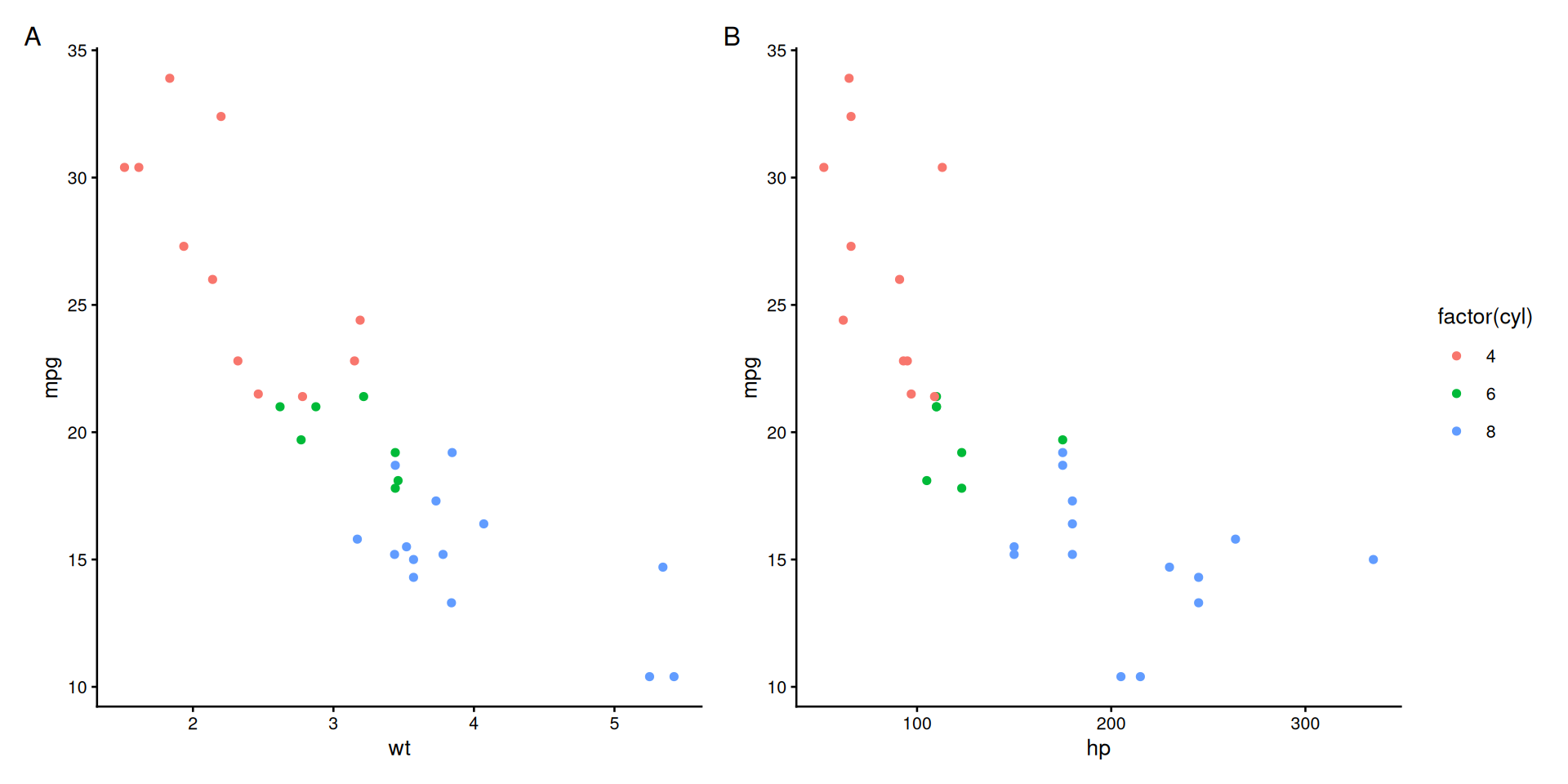
Shared Axes with plot_layout()
- Default behavior: Each plot keeps its own axes
collect_x: Remove duplicate x-axes when plots are stacked vertically (same x-scale)collect_y: Remove duplicate y-axes when plots are side-by-side (same y-scale)collect: Remove both x and y axes (same scales in both directions)- Apply to groups where it makes sense before combining
Saving patchwork Figures
# Create combined figure
combined <- (p1 | p2) / (p3 | p4) +
plot_annotation(tag_levels = 'A')
# Save as vector (recommended)
ggsave("figure1.svg", combined, width = 10, height = 8)
ggsave("figure1.pdf", combined, width = 10, height = 8)
# Save as high-res raster if needed
ggsave("figure1.png", combined, width = 10, height = 8, dpi = 300)patchwork vs Manual Editing: Decision Guide
patchwork (in R)
✅ Fully reproducible
✅ Easy to update
✅ Automatic alignment
✅ Consistent styling
✅ Version controlled
⚠️ Less layout flexibility
Use patchwork when:
- All panels are ggplot2 objects
- Need reproducible figures
- Figures may need updates
- Sharing code with collaborators
Inkscape (manual editing)
✅ Pixel-perfect control
✅ Mix with non-R content
✅ Complex annotations
❌ Not reproducible
❌ Manual re-editing
❌ Easy to break
Use Inkscape when:
- Need pixel-perfect alignment
- Adding photos, diagrams, or non-R content
- Very complex custom layouts
- Final publication polish only
PowerPoint Import
The Copy-Paste Problem
What happens when you copy from RStudio:
- Pastes as low-resolution bitmap
- Looks OK on screen (72 DPI)
- Terrible when projected
- Pixelated and blurry
- Doesn’t scale well
- The “Copy Plot to Clipboard” → “Copy as Metafile” corrupts plot symbols. Save as SVG instead.
The Solution: Save First, Insert Second
Never copy-paste!
Instead:
- Save plot as file
- Insert file into PowerPoint
- Maintain quality
Vector Formats for PowerPoint
Three options:
SVG: Good support in modern PowerPoint → Use this!
- Editable after import
- Preserves vector format
- Exports as SVG/PDF maintain vector quality
EMF: Windows only, obsolete and of no benefit in newer PowerPoint. Requires
devEMFPDF: Very poorly supported! Low resolution import (rasterized!) and no editing
PNG: If you must use raster, then 300+ DPI
SVG Editing in PowerPoint
SVG is Editable in PowerPoint
Modern PowerPoint supports SVG editing:
- Insert SVG file into PowerPoint
- Right-click → Ungroup (or Convert to Shape)
- Individual elements become editable
- Modify colors, text, positions
- Compose multi-panel figures
- Export as SVG/PDF to preserve vector format
PowerPoint can be your figure composition tool!
Ungrouping May Break Complex SVGs
Be careful when ungrouping:
- Complex SVGs may lose gradients, patterns, or effects
- Some plot elements might break apart unexpectedly
- Text rendering may change
- Clipping paths may be lost
Recommendation:
- Keep a backup copy before ungrouping
- Test with your specific plots first
- For complex figures, consider Inkscape instead if editing is needed.
Factor Ordering
The Problem: Alphabetical Ordering
# Create data with logical categories
category_df <- data.frame(
category = c("Low", "Medium", "High", "Very High", "Low", "High"),
value = c(10, 20, 15, 30, 12, 25)
)
# R defaults to alphabetical!
p1 <- ggplot(category_df, aes(x = category, y = value)) +
geom_col(fill = "coral") +
theme_classic(base_size = 12) +
labs(title = "Alphabetical (Wrong!)")# Specify levels explicitly
category_df$category <- factor(category_df$category,
levels = c("Low", "Medium", "High", "Very High"))
# Now plots use logical order!
p2 <- ggplot(category_df, aes(x = category, y = value)) +
geom_col(fill = "steelblue") +
theme_classic(base_size = 12) +
labs(title = "Logical Order (Correct!)")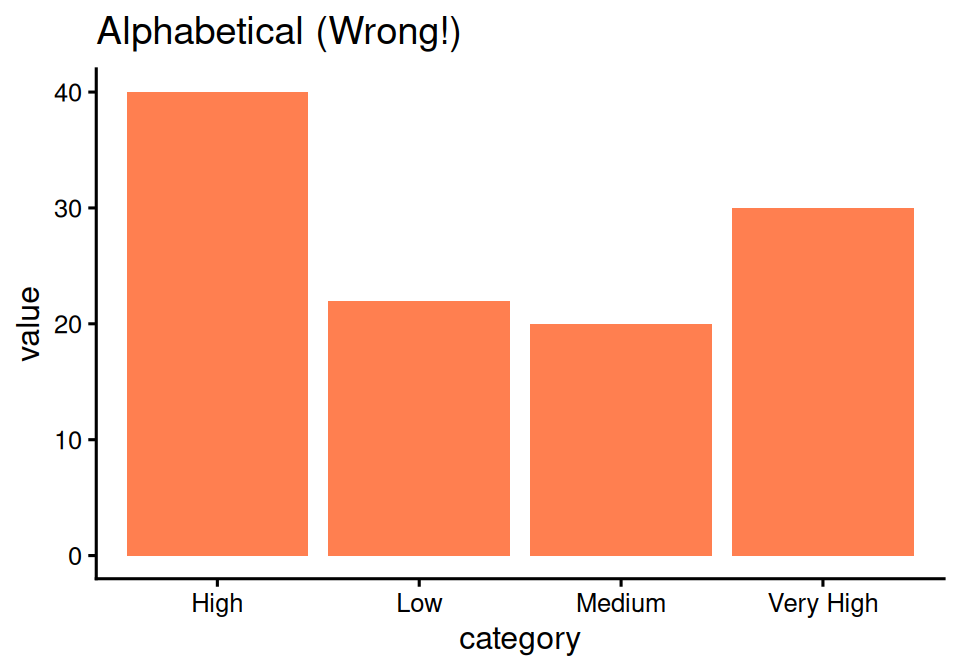
Random order makes no sense!
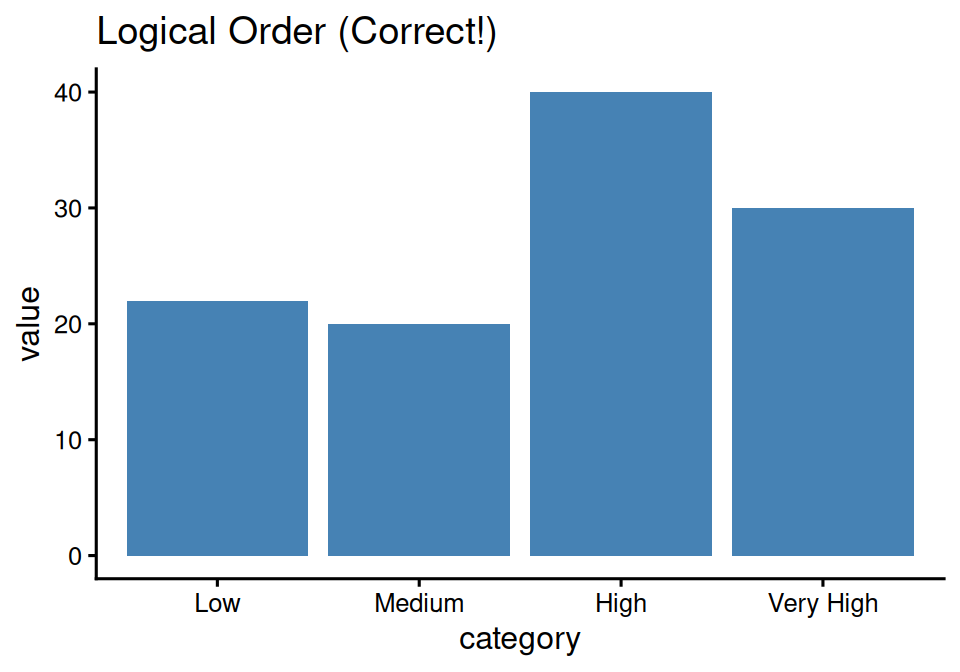
Much better - order makes sense!
Solution: forcats Package
Part of tidyverse, designed for factor manipulation
fct_reorder(): Order by Another Variable
# Order diamond cuts by mean price
p <- diamonds %>%
group_by(cut) %>%
summarise(mean_price = mean(price)) %>%
ggplot(aes(x = fct_reorder(cut, mean_price),
y = mean_price,
fill = cut)) +
geom_col(show.legend = FALSE) +
coord_flip() +
labs(x = "Diamond Cut",
y = "Mean Price ($)",
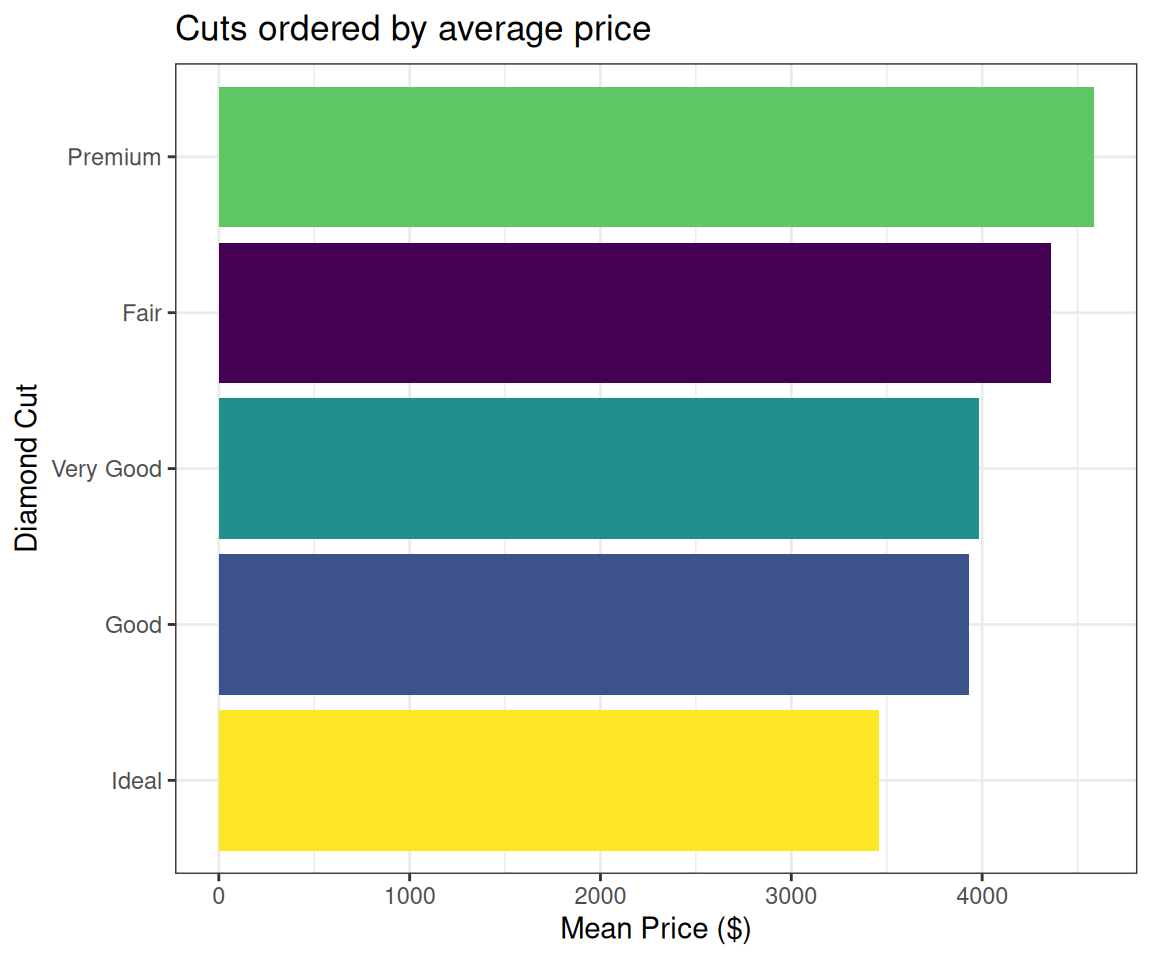
title = "Cuts ordered by average price")Bars ordered by length!
fct_infreq(): Order by Frequency
fct_inorder(): Order by Appearance
# Create data with specific order
treatment_data <- data.frame(
treatment = c("Control", "Low Dose",
"Medium Dose", "High Dose",
"Control", "Low Dose",
"Medium Dose", "High Dose"),
response = c(10, 12, 15, 18,
11, 13, 16, 19)
)
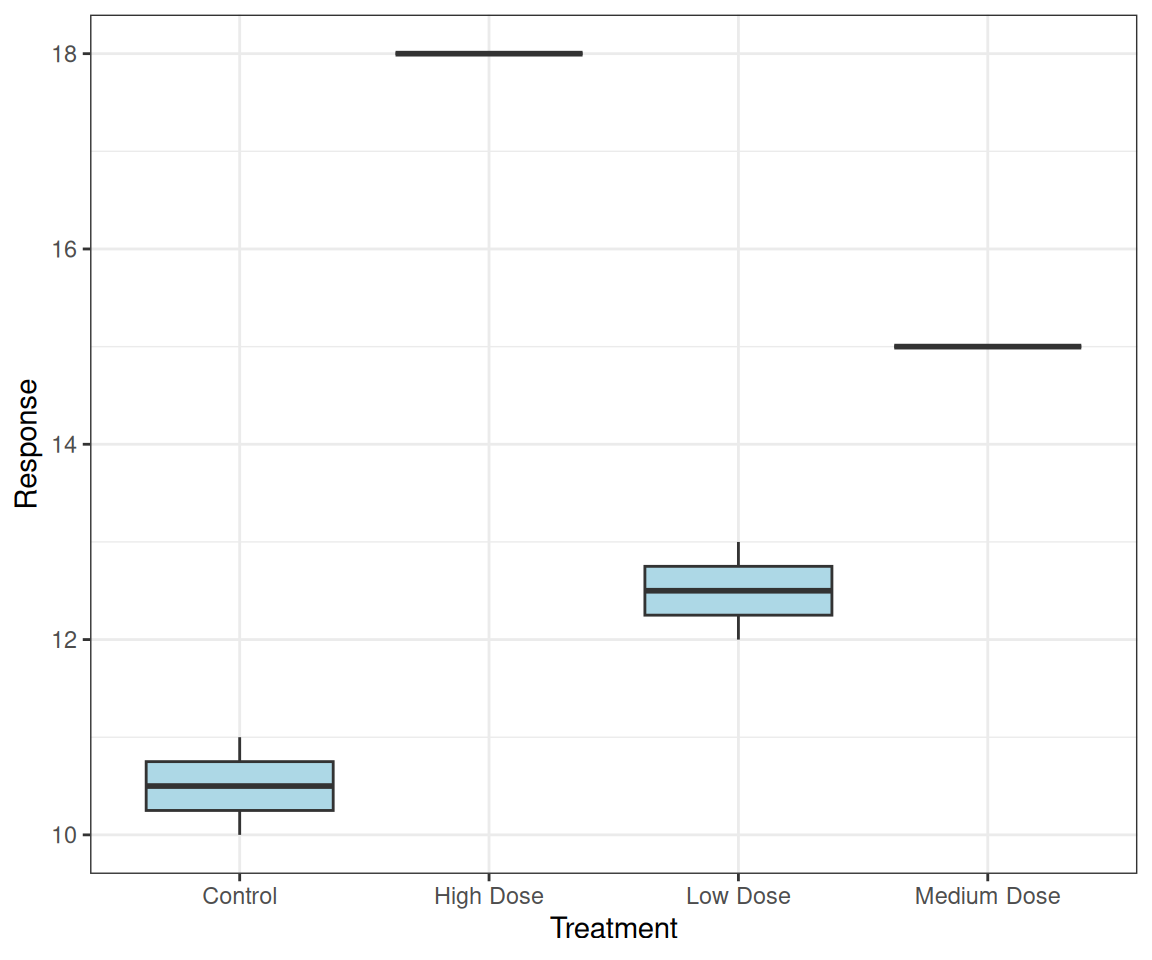
# Keep order as they appear in data
p <- ggplot(treatment_data,
aes(x = fct_inorder(treatment),
y = response)) +
geom_boxplot(fill = "lightblue") +
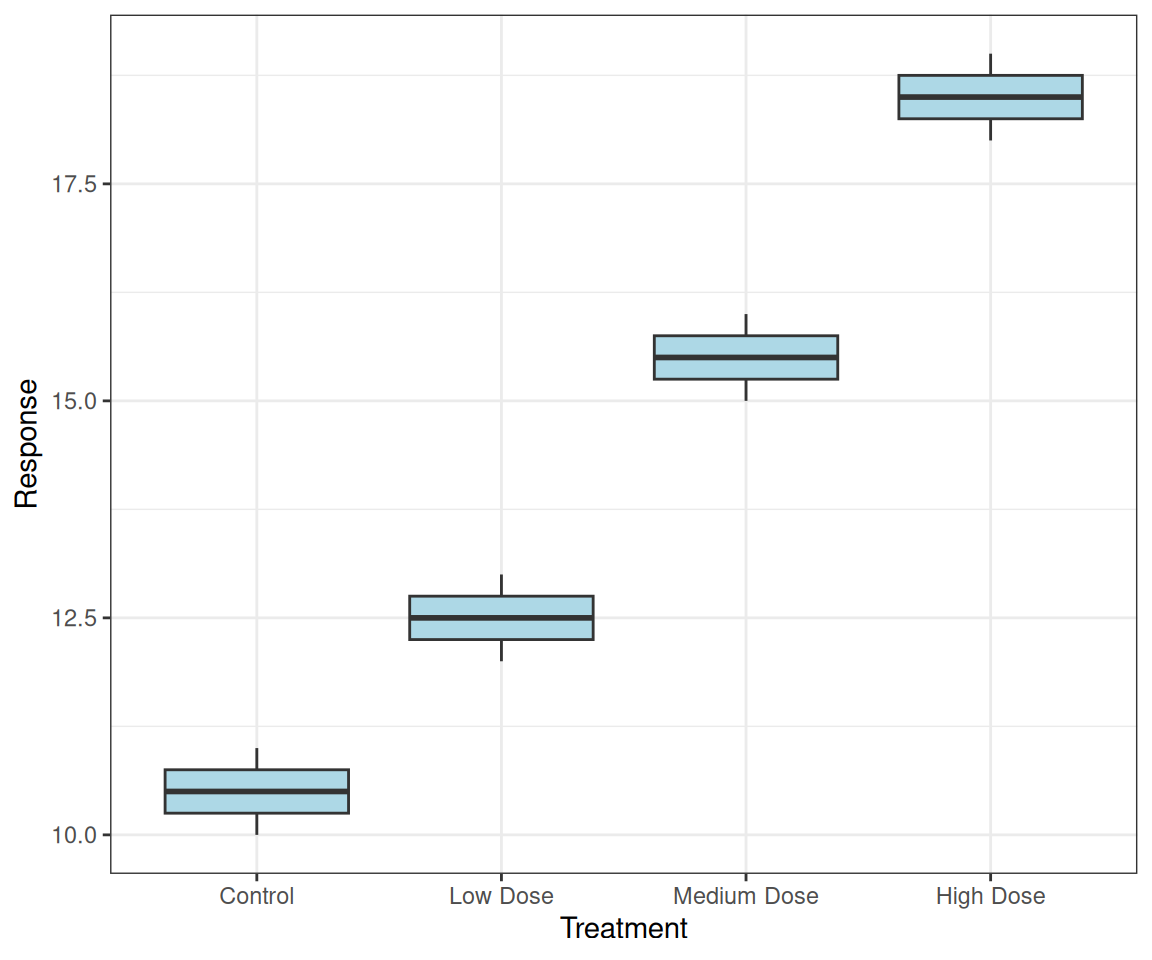
labs(x = "Treatment", y = "Response")Preserves the order from your data!
Natural Sorting: var1, var2, …, var10
The problem with alphabetical sorting:
# Create data with numbered variables
var_data <- data.frame(
variable = rep(c("var1", "var2", "var10", "var20"), each = 5),
value = rnorm(20, mean = rep(c(10, 15, 20, 25), each = 5), sd = 2)
)
# Alphabetical order: var1, var10, var2, var20 (wrong!)
p <- ggplot(var_data, aes(x = variable, y = value)) +
geom_boxplot(fill = "coral") +
labs(title = "Alphabetical: var1, var10, var2, var20")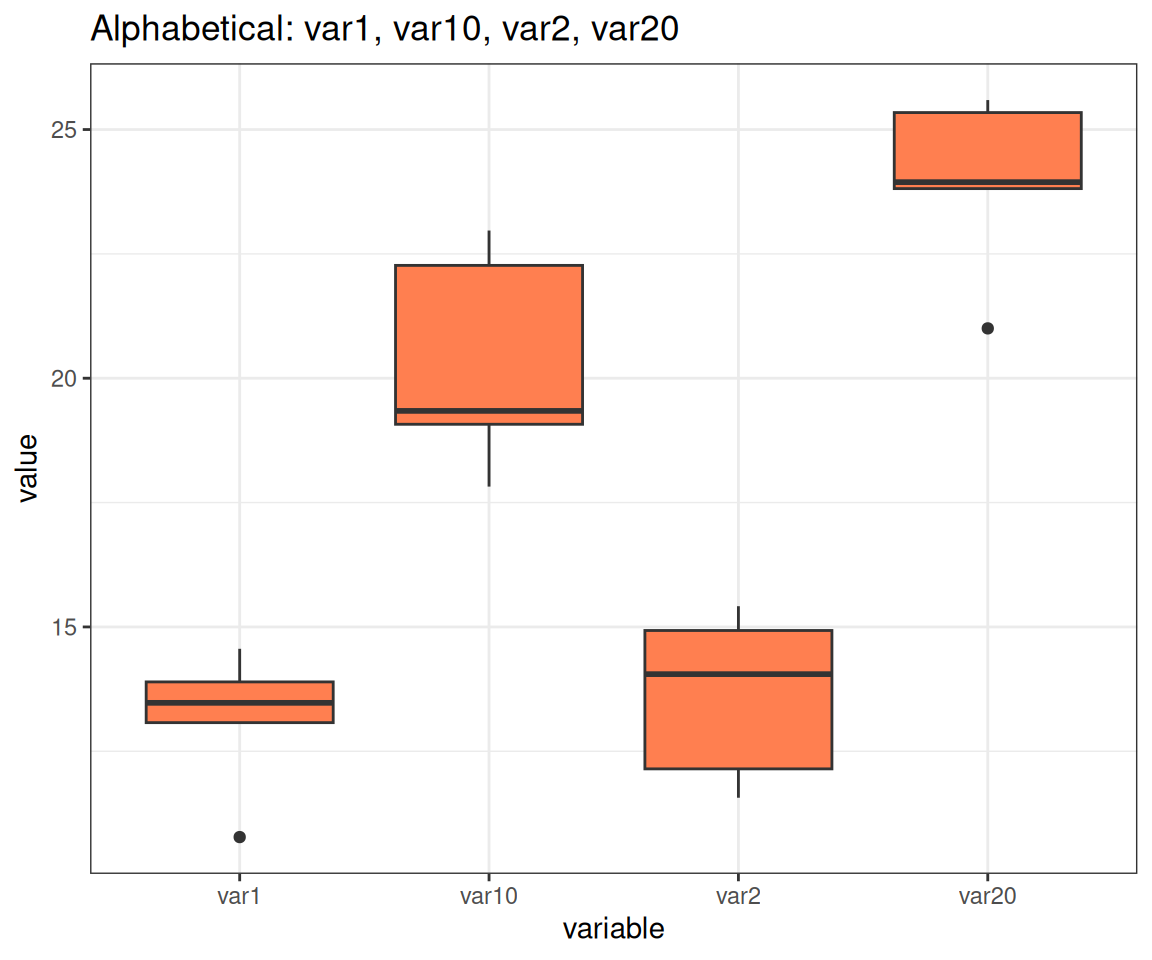
Natural Sorting: The Solution
# Use gtools::mixedsort() for natural/alphanumeric sorting
library(gtools)
var_data$variable <- factor(var_data$variable,
levels = mixedsort(unique(var_data$variable)))
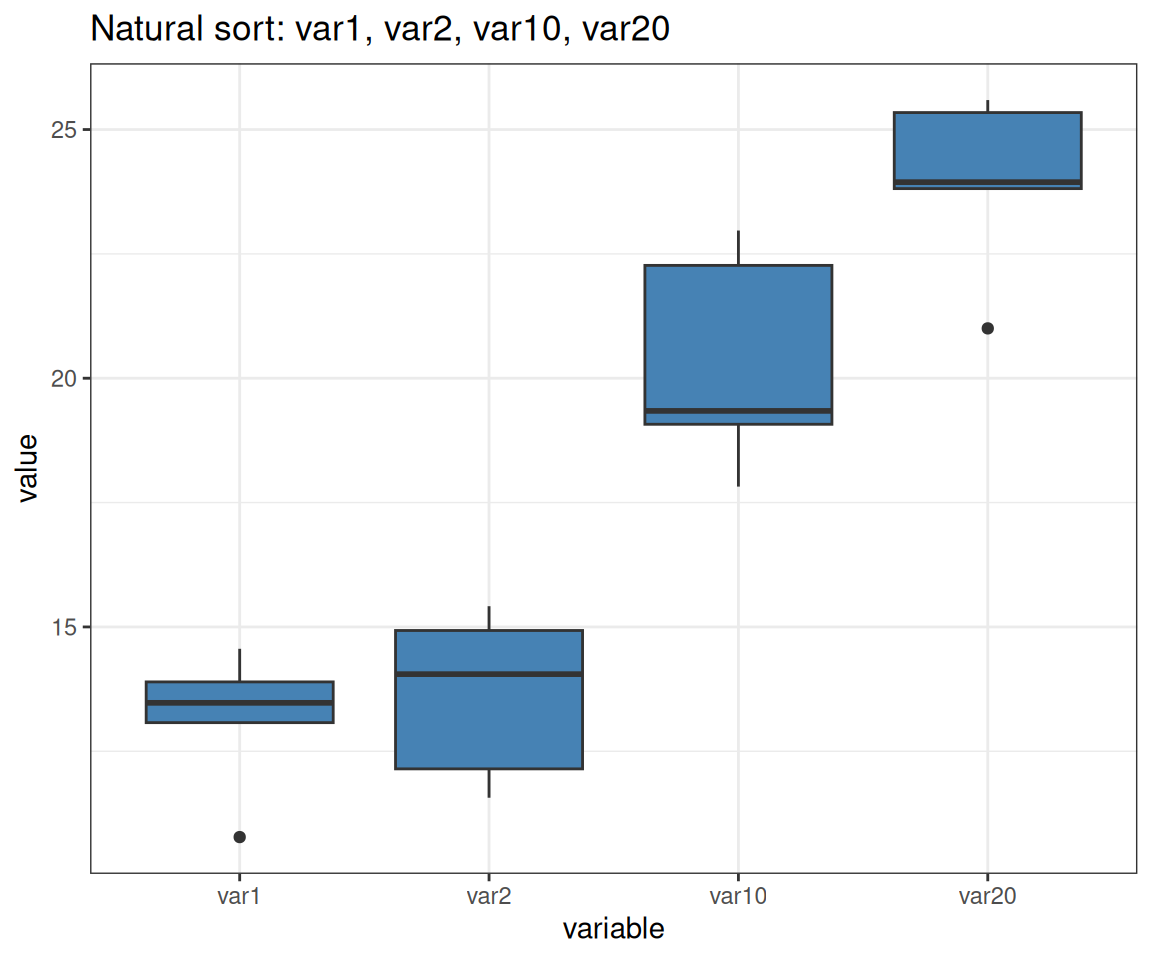
# Natural order: var1, var2, var10, var20 (correct!)
p <- ggplot(var_data, aes(x = variable, y = value)) +
geom_boxplot(fill = "steelblue") +
labs(title = "Natural sort: var1, var2, var10, var20")Note: forcats::fct_inseq() only works if factor levels are purely numeric strings (e.g., “1”, “2”, “10”), not mixed alphanumeric like “var1”, “var10”
fct_rev(): Reverse Order
fct_relevel(): Move Specific Levels
# Move "Control" to front for treatment groups
treatment_data <- data.frame(
treatment = c("Low Dose", "High Dose", "Control",
"Medium Dose", "Low Dose", "Control"),
response = c(12, 18, 10, 15, 13, 11)
)
p <- treatment_data %>%
mutate(treatment = fct_relevel(treatment, "Control")) %>%
ggplot(aes(treatment, response)) +
geom_boxplot(fill = "lightblue") +
labs(x = "Treatment", y = "Response")Control always shown first
Common in experimental data!
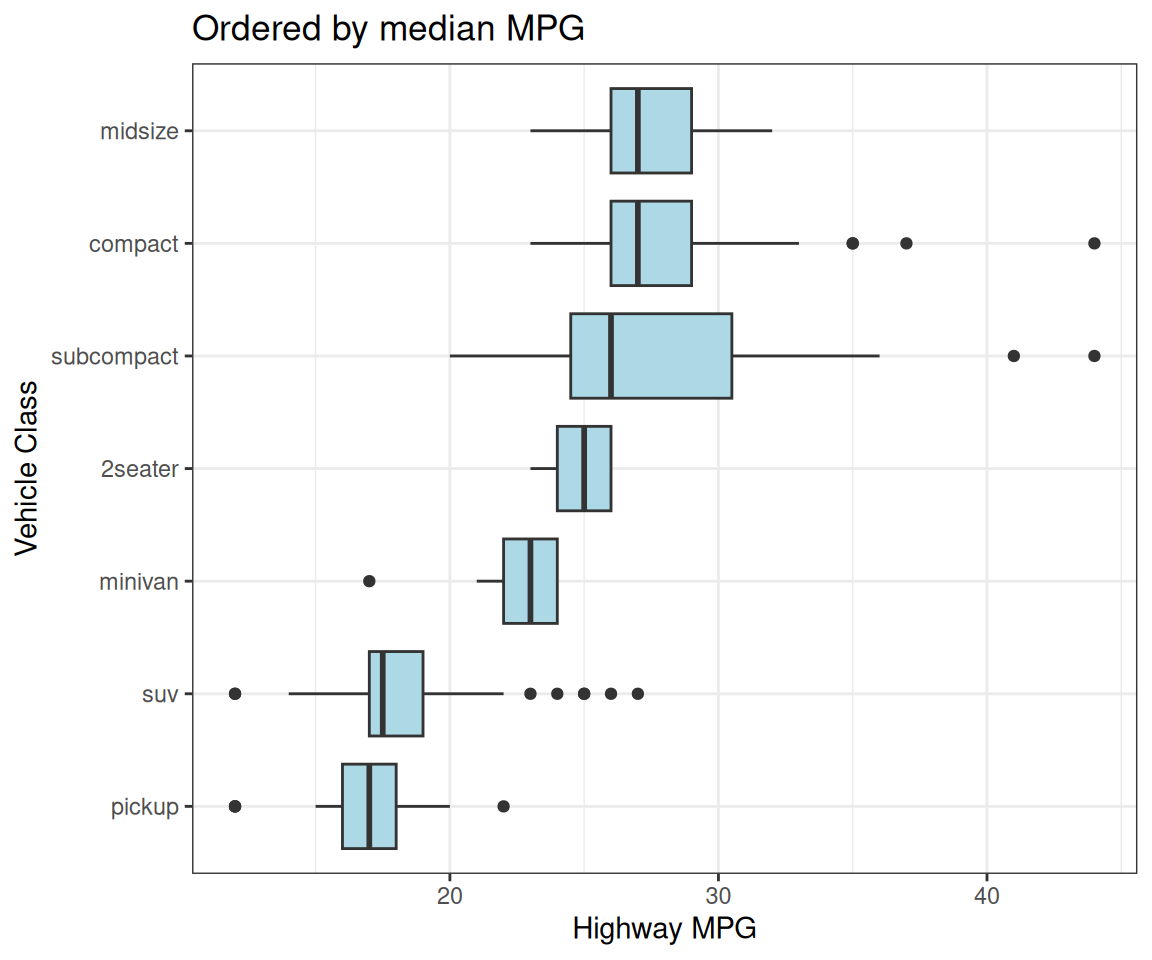
Facet Ordering
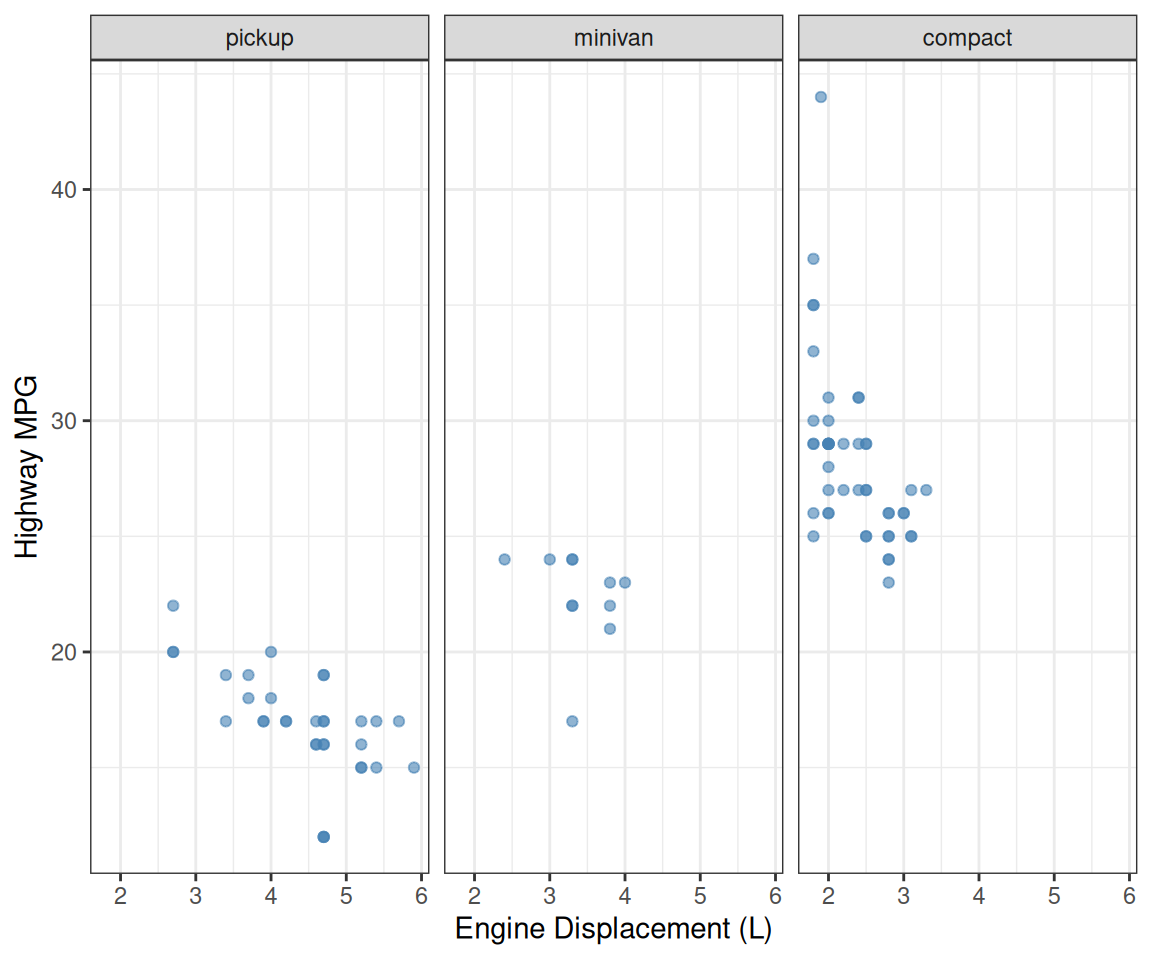
# Order facets by median highway mpg for each vehicle class
p <- mpg %>%
filter(class %in% c("pickup", "minivan", "compact")) %>%
mutate(class = fct_reorder(class, hwy, median)) %>%
ggplot(aes(x = displ, y = hwy)) +
geom_point(color = "steelblue", alpha = 0.6) +
facet_wrap(~class) +
labs(x = "Engine Displacement (L)", y = "Highway MPG")Facet panels in meaningful order!
⚠️ The Danger of Numeric Factors
Converting between factor and numeric can destroy your data!
The danger: Many functions silently convert factors to integers!
⚠️ Missing Levels: The Silent Data Corruption
Even worse: missing levels get renumbered!
Returns: 1 2 3 4 1 2 3 4
Your subject 4 became 3!
Your subject 5 became 4!
This is catastrophic for analysis!
Your statistical models and plots will use the wrong subject numbers. Always use as.numeric(as.character(factor)) not as.numeric(factor).
Or better yet. NEVER use numbers for categorical data!
⚠️ Numeric Factors After Reordering
# A tibble: 5 × 5
dose response dose_ordered wrong correct
<fct> <dbl> <fct> <dbl> <dbl>
1 0 5 0 1 0
2 10 25 10 2 10
3 50 80 50 5 50
4 100 70 100 4 100
5 200 30 200 3 200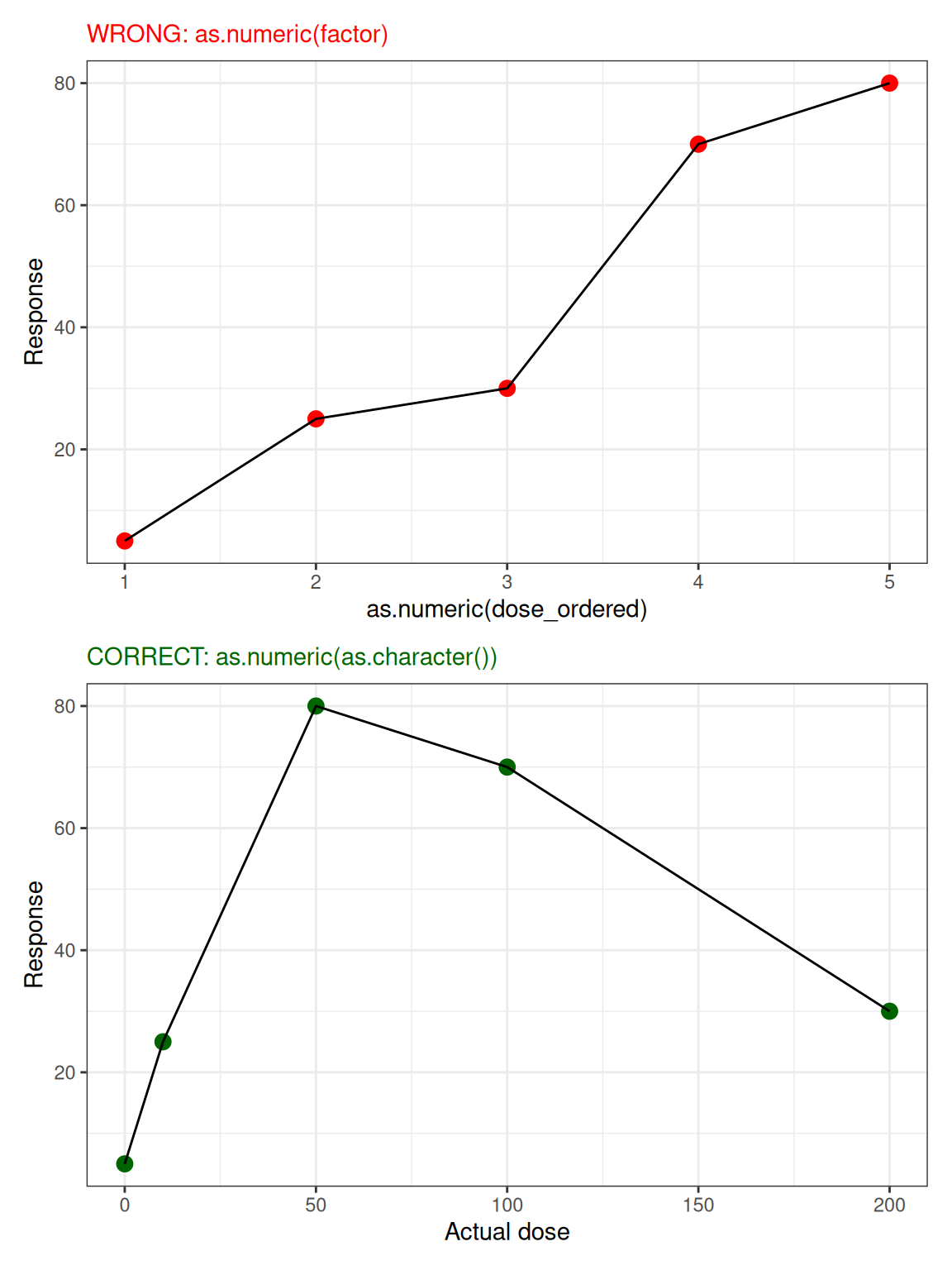
Recommendations: Avoid Numeric Factors
Best Practices
- Never store numeric values as factors unless they represent categories
- Prefix categorical numbers: Use “Group_1”, “Group_2” instead of “1”, “2”
- Check imported data: CSV imports often convert numbers to factors
- Use
readr::read_csv()instead ofread.csv()- better type detection and no implicit conversion to factors. - Always convert via character:
as.numeric(as.character(factor))notas.numeric(factor) - Check with
str()before analysis to verify data types
Bad: Numeric categories
forcats Cheat Sheet
library(forcats)
fct_reorder(f, x, fun) # Order by another variable
fct_infreq(f) # Order by frequency
fct_inorder(f) # Order by appearance in data
fct_inseq(f) # Order by numeric value (if purely numeric)
fct_rev(f) # Reverse current order
fct_relevel(f, "A", "B") # Move specific levels to front
fct_recode(f, new = "old") # Rename levels
fct_lump_n(f, n = 5) # Keep top n, lump others as "Other"
fct_explicit_na(f) # Make NA a visible level
# For natural sort (var1, var2, var10):
factor(x, levels = gtools::mixedsort(unique(x)))Debugging Factor Issues
# Example: vehicle classes in mpg dataset
vehicle_class <- factor(mpg$class)
# Check level order
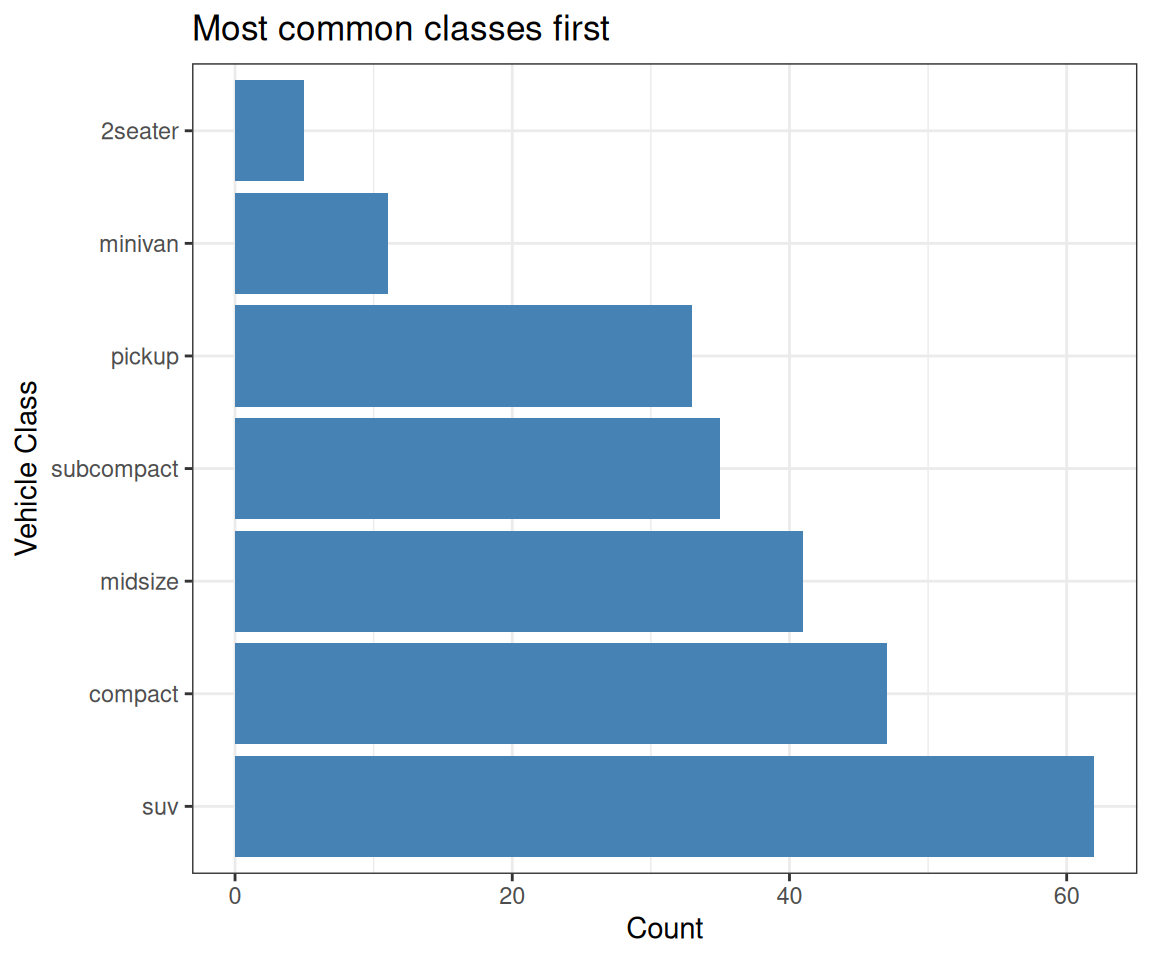
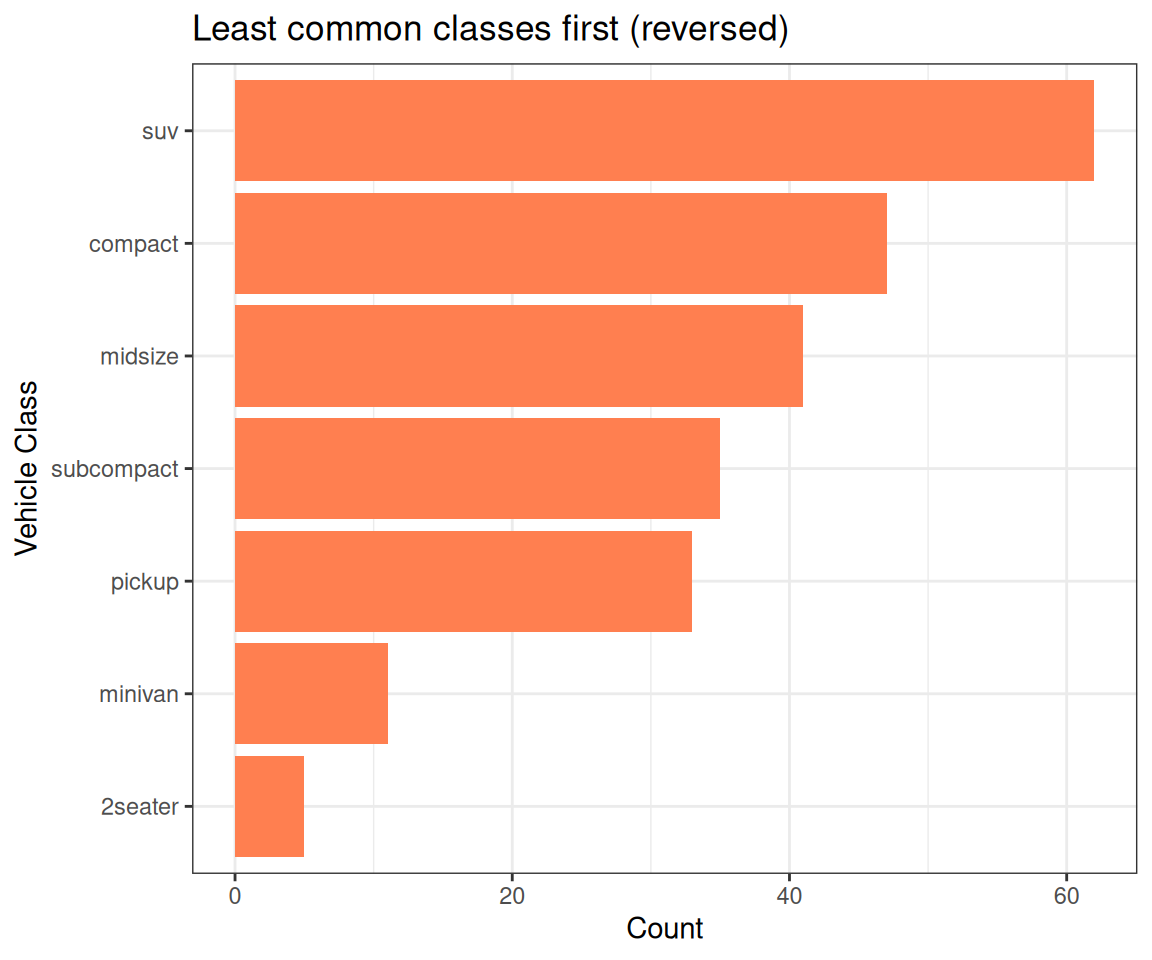
vehicle_class %>% levels()[1] "2seater" "compact" "midsize" "minivan" "pickup"
[6] "subcompact" "suv" Factor w/ 7 levels "2seater","compact",..: 2 2 2 2 2 2 2 2 2 2 ...vehicle_class
2seater compact midsize minivan pickup subcompact suv
5 47 41 11 33 35 62 Key Takeaways & Best Practices
Remember
The Problem:
- R defaults to alphabetical factor ordering - usually wrong!
- Numeric factors are dangerous - converting destroys your data
- Missing levels get renumbered - group 4 becomes 3!
The Solutions:
- forcats package provides powerful ordering tools:
fct_reorder()orders by another variable (most useful!)fct_infreq()orders by frequencyfct_relevel()to put control/baseline firstfct_rev()to reverse order
- Manual levels for logical ordering (Low/Med/High, months, etc.)
- Prefix categorical numbers: “Group_1” not “1”
Always:
- Check factor order before plotting with
str()orlevels() - Convert via character:
as.numeric(as.character(f))notas.numeric(f) - Think about your reader - what order makes sense?
Interactive Plots
The Appeal of Interactivity
Why use interactive plots?
- 🖱️ Hover to see exact values
- 🔍 Zoom and pan
- 👁️ Toggle traces on/off
- 📊 Great for exploration
- 🎯 Excellent for presentations
Basic ggplotly Example
library(ggplot2)
library(plotly)
# Create clean data for tooltips
mtcars_clean <- mtcars
mtcars_clean$Cylinders <- factor(mtcars$cyl)
# Create ggplot
p <- ggplot(mtcars_clean, aes(wt, mpg, color = Cylinders)) +
geom_point(size = 3) +
geom_smooth(method = "lm", se = FALSE) +
scale_color_brewer(palette = "Set1") +
theme_minimal() +
labs(title = "Fuel Efficiency by Weight",
x = "Weight (1000 lbs)",
y = "Miles per Gallon",
color = "Cylinders")Try hovering over points!
- See exact values
- Toggle series on/off
- Zoom and pan
Customizing Tooltips
Now hover shows car name and horsepower!
Use the text aesthetic and tooltip parameter to customize what appears on hover.
The Saving Problem
Challenge
Interactive plots are HTML widgets, not static images!
Can’t just save as PDF or PNG traditionally.
Solution: Save as HTML (Keep Interactivity)
About selfcontained parameter:
selfcontained = TRUE: Bundles all JavaScript/CSS into one file- Perfect for emailing or sharing
- No external dependencies needed
- Larger file size
selfcontained = FALSE: Creates separate library files- Smaller main HTML file
- Requires folder structure to be maintained
Uses:
- Email the HTML file directly
- Upload to website
- Opens in any web browser!
Quarto/Markdown Integration
In Quarto, ggplotly works seamlessly:
Perfect for modern scientific reports!
3D Plots
Linked Plots with Crosstalk
# Create shared data
mtcars$car_name <- rownames(mtcars)
shared_data <- SharedData$new(mtcars, ~car_name)
# Create linked plots
p1 <- plot_ly(shared_data, x = ~wt, y = ~mpg, type = 'scatter', mode = 'markers')
p2 <- plot_ly(shared_data, x = ~hp, y = ~mpg, type = 'scatter', mode = 'markers')
# Display with filter on top
bscols(widths = 12, filter_checkbox("cyl", "Cylinders:", shared_data, ~cyl, inline = TRUE))
subplot(p1, p2, nrows = 1, shareY = TRUE)Hybrid Approach: Both Versions
# Create output directory if needed
dir.create("plots/10_interactive", recursive = TRUE, showWarnings = FALSE)
# Create base plot
p_static <- ggplot(mtcars, aes(wt, mpg, color = factor(cyl))) +
geom_point(size = 3) +
theme_classic(base_size = 12)
# Static version for publication
ggsave("plots/10_interactive/figure1.pdf", p_static, width = 7, height = 5)
# Interactive version for supplement/website
p_interactive <- ggplotly(p_static)
saveWidget(p_interactive, "plots/10_interactive/figure1_interactive.html")
# Best of both worlds!Sharing Interactive Plots
Options:
- Email HTML file (if selfcontained = TRUE)
- Upload to web server
- GitHub Pages (free hosting)
- Shiny app (for more complex interactions)
Limitations of ggplotly
Not all ggplot2 features convert:
- Some geoms don’t translate well
- Complex annotations may be lost
- Custom themes partially supported
- Facets work but can be slow
Test your conversion!
Alternative: ggiraph
library(ggiraph)
# Create plot with interactive elements
mtcars$car_name <- rownames(mtcars)
# Create rich tooltips with HTML formatting
mtcars$tooltip_text <- paste0(
"<b>", mtcars$car_name, "</b><br>",
"Weight: ", round(mtcars$wt, 2), " (1000 lbs)<br>",
"MPG: ", mtcars$mpg, "<br>",
"HP: ", mtcars$hp, "<br>",
"Cylinders: ", mtcars$cyl
)
p <- ggplot(mtcars, aes(wt, mpg,
tooltip = tooltip_text,
data_id = car_name)) +
geom_point_interactive(aes(color = factor(cyl)), size = 3) +
theme_minimal()Key Takeaways
Best Practices & Summary
Core concepts:
- Interactive plots are HTML widgets, not images
- Use
ggplotly()to convert ggplot2 plots instantly - Use
saveWidget()withselfcontained = TRUEto save
When to use what:
- Interactive: exploration, presentations, HTML reports
- Static: publications, print, PDF reports
Tips:
- Create both versions when possible
- Sample large datasets to keep HTML file manageable
- Test thoroughly - not all ggplot2 features convert
- Consider ggiraph if ggplotly doesn’t preserve your styling
- Use selfcontained = TRUE for easy sharing
References
Key Papers and Books
Key Papers
Essential research on color use, perception, and data visualization:
- Crameri, Shephard, and Heron (2020) - The misuse of color in science communication
- Borkin et al. (2011) - Evaluation of visualization effectiveness
- Gołębiowska and Çöltekin (2022) - Problems with rainbow color schemes
- Heron, Crameri, and Shephard (2021) - Rainbow colormaps and their issues
Books
- Wilke (2019) - Fundamentals of Data Visualization - Comprehensive guide to visualization principles (free online)
- R Graphics Cookbook by Winston Chang - Practical recipes for ggplot2 (free online)
Color Resources
Tools
- ColorBrewer - Interactive tool for selecting color palettes for maps and data visualization
- Coblis - Color Blindness Simulator - Upload images to see how they appear with different types of color vision deficiency
R Packages
- viridis - Perceptually uniform color scales - https://cran.r-project.org/package=viridis
- RColorBrewer - ColorBrewer palettes for R
- colorspace - Color manipulation and assessment
- dichromat - Simulate color blindness
- paletteer - Collection of 2000+ palettes
Getting help
Documentation
- ggplot2 Reference - Complete function reference
- plotly for R - Interactive plotting library
Community
- Stack Overflow - Q&A for ggplot2 questions
Software
Essential Tools
- R - The R programming language
- RStudio - Integrated development environment
- Quarto - Scientific and technical publishing system
Helpful Extensions
- esquisse - Interactive ggplot2 builder
- ggThemeAssist - Visual theme customization
Inspiration
Outstanding Examples
- BBC Visual and Data Journalism - BBC’s ggplot2 cookbook
- Financial Times Visual Vocabulary - Guide to chart types
Related Presentations
- This presentation: https://stanstrup.github.io/figure_presentation/
- Heatmaps guide: https://stanstrup.github.io/heatmaps.html
Contributing
Found a useful resource not listed here? Contributions are welcome!
- GitHub: https://github.com/stanstrup/figure_presentation
- Issues: Report problems or suggest additions
- Pull Requests: Contribute improvements
Citation
If you find this guide useful in your work, please cite:
Stanstrup, J. (2025). Academic Figures: Common Pitfalls and Best Practices.
https://stanstrup.github.io/figure_presentation/Academic Figure Pitfalls